Beruflich Dokumente
Kultur Dokumente
Course Hero
Hochgeladen von
fbihansip0%(1)0% fanden dieses Dokument nützlich (1 Abstimmung)
18K Ansichten3 SeitenThis document reports the results of a chi-square test performed on progeny data from a genetic cross. The observed and expected ratios across four phenotypes (PL, Pl, pL, pl) are shown, and a chi-square value of 134.74 is calculated, exceeding the critical value for 3 degrees of freedom and rejecting the hypothesis that the alleles segregate independently. Further calculations estimate the recombination frequency at 20% and show the expected ratios assuming linkage. A second chi-square test is reported with the expected ratios adjusted for linkage, but the hypothesis of linkage is still rejected at the 5% significance level.
Originalbeschreibung:
ini greget bro
Copyright
© © All Rights Reserved
Verfügbare Formate
DOCX, PDF, TXT oder online auf Scribd lesen
Dieses Dokument teilen
Dokument teilen oder einbetten
Stufen Sie dieses Dokument als nützlich ein?
Sind diese Inhalte unangemessen?
Dieses Dokument meldenThis document reports the results of a chi-square test performed on progeny data from a genetic cross. The observed and expected ratios across four phenotypes (PL, Pl, pL, pl) are shown, and a chi-square value of 134.74 is calculated, exceeding the critical value for 3 degrees of freedom and rejecting the hypothesis that the alleles segregate independently. Further calculations estimate the recombination frequency at 20% and show the expected ratios assuming linkage. A second chi-square test is reported with the expected ratios adjusted for linkage, but the hypothesis of linkage is still rejected at the 5% significance level.
Copyright:
© All Rights Reserved
Verfügbare Formate
Als DOCX, PDF, TXT herunterladen oder online auf Scribd lesen
0%(1)0% fanden dieses Dokument nützlich (1 Abstimmung)
18K Ansichten3 SeitenCourse Hero
Hochgeladen von
fbihansipThis document reports the results of a chi-square test performed on progeny data from a genetic cross. The observed and expected ratios across four phenotypes (PL, Pl, pL, pl) are shown, and a chi-square value of 134.74 is calculated, exceeding the critical value for 3 degrees of freedom and rejecting the hypothesis that the alleles segregate independently. Further calculations estimate the recombination frequency at 20% and show the expected ratios assuming linkage. A second chi-square test is reported with the expected ratios adjusted for linkage, but the hypothesis of linkage is still rejected at the 5% significance level.
Copyright:
© All Rights Reserved
Verfügbare Formate
Als DOCX, PDF, TXT herunterladen oder online auf Scribd lesen
Sie sind auf Seite 1von 3
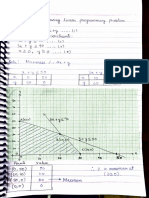
phenotype PL Pl pL Pl
Observe (O) 284 21 21 55
Expected ratio 9/16 3/16 3/16 1/16
Expected (E) 214,3125 71,4375 71,4375 23,8125
Expected is from = expected ratio X total progeny observed
Total number of progeny is 381 individuals.
Chi-square formula is X
2
= {(o-e)
2
}/e
Phenotype PL Pl pL pl
O-E 284-214,3125 21-71,4375 21-71,4375 55-23,8125
69,6875 -50,4375 -50,4375 31,1875
(O-E)
2
4856,347656 2543,941406 2543,941406 972,660
(O-E)
2
/ E 22,67 35,61 35,61 40,85
(O-E)
2
/ E 134,74
DF= phenotype-1
DF= 4-1 = 3
Look to the r= 3 (df)
And = 5% or 0,05 (in 0,950 row)
The value is 7,815, because the X
2
counting > X
2
table
So the hypothesis that the alel segragate freely(independently) and all equaly viable are rejected.
How about linkage???
Notice That homozygote recessive only can be inherited If Two pl genotype are met.
So 55/341= 0,16~
So if probabilitas of ppll is probabilitas of pl times pl, so if the ppll is 0,16 then the pl is the square
root of 0,16, then pl is 0,4.
Next pl is one of the parental genotype beside PL, so the sum of parental is (PL+pl) is 0,4+0,4= 0,8....
Then the recombinan is 0,2 or 20%.
So the distance is 20 mU.
PL (0,4) Pl (0,1) pL (0,1) pl (0,4)
PL (0,4) PPLL (0,16) PPLl (0,04) PpLL (0,04) PpLl (0,16)
Pl (0,1) PPLl (0,04) PPll (0,01) PpLl (0,01) Ppll (0,04)
pL (0,1) PpLL (0,04) PpLl (0,01) ppLL (0,01) ppLl (0,04)
pl (0,4) PpLl (0,16) Ppll (0,04) ppLl (0,04) Ppll (0,16)
Expected is from = expected ratio X total progeny observed
phenotype PL Pl pL Pl
Observe (O) 284 21 21 55
Expected ratio 0,66 0,09 0,09 0,16
Expected (E) 251,46 34,29 34,29 60,96
O-E 32,54 -13,29 -13,29 -5,96
(O-E)
2
1058,8516 176,6241 176,6241 35,5216
(O-E)
2
/ E 4,21 5,15 5,15 0,42
(O-E)
2
/ E 14,93
DF= phenotype-1
DF= 4-1 = 3
Look to the r= 3 (df)
And = 5% or 0,05 (in 0,950 row)
The value is 7,815, because the X
2
counting > X
2
table
Still the hypotesis for lingkage is rejected.
If compared, from linked or not, it is more likely to be linked.... but still not well accepted in 0,05.
Das könnte Ihnen auch gefallen
- Categorical Data Analysis Selected Solutions by AgrestiDokument30 SeitenCategorical Data Analysis Selected Solutions by AgrestiNeil Bot67% (3)
- Modern Digital and Analog Communications Systems - B P Lathi Solutions ManualDokument155 SeitenModern Digital and Analog Communications Systems - B P Lathi Solutions Manualsandy_00991% (88)
- Papoulis Solutions ManualDokument186 SeitenPapoulis Solutions ManualSoroushGooran50% (2)
- Question Bank With AnswersDokument165 SeitenQuestion Bank With Answersvignanaraj50% (2)
- On The Calculus of Smarandache FunctionDokument8 SeitenOn The Calculus of Smarandache FunctionMia AmaliaNoch keine Bewertungen
- Math - McGraw Hil - Probability Random Variables and Stochastic Processes Solutions Manual - Papoulis - 2002Dokument187 SeitenMath - McGraw Hil - Probability Random Variables and Stochastic Processes Solutions Manual - Papoulis - 2002Mustafa DinçNoch keine Bewertungen
- Profile Likelihood: N X X XDokument4 SeitenProfile Likelihood: N X X XjuntujuntuNoch keine Bewertungen
- APA Post Mid TermDokument12 SeitenAPA Post Mid Termabhinavmalviya077188Noch keine Bewertungen
- Comparison of ND Scattering Calculations Using Different Representations of Realistic PotentialsDokument16 SeitenComparison of ND Scattering Calculations Using Different Representations of Realistic PotentialsWolfgang SchadowNoch keine Bewertungen
- 12 Maths Exemplar Chapter 13 PDFDokument29 Seiten12 Maths Exemplar Chapter 13 PDFMohammed IrshadNoch keine Bewertungen
- Solutions To ExercisesDokument54 SeitenSolutions To ExercisesNelsonNoch keine Bewertungen
- Journal of Engineering Physics and Thermophysics, Vol. 72, No. 3, 1999Dokument16 SeitenJournal of Engineering Physics and Thermophysics, Vol. 72, No. 3, 1999thotalnNoch keine Bewertungen
- R C R CC R + (H) : Delay Differential EquationsDokument14 SeitenR C R CC R + (H) : Delay Differential Equationsrifky fauziNoch keine Bewertungen
- J.N. Murrell Et Al - Potential Energy Curves of The Lower States of CN +Dokument6 SeitenJ.N. Murrell Et Al - Potential Energy Curves of The Lower States of CN +MaxnamewNoch keine Bewertungen
- Probability NotesDokument18 SeitenProbability NotesVishnuNoch keine Bewertungen
- Chapter 1 Intro and Basic Concepts: Nominal: Cannot Be Ordered Ordinal: Can Be OrderedDokument40 SeitenChapter 1 Intro and Basic Concepts: Nominal: Cannot Be Ordered Ordinal: Can Be OrderedMartijnKruijzenNoch keine Bewertungen
- Leep 213Dokument29 SeitenLeep 213Koyal GuptaNoch keine Bewertungen
- Calculation of The Nonlinear Optical Properties Molecules: P, F P Po F - FDokument6 SeitenCalculation of The Nonlinear Optical Properties Molecules: P, F P Po F - FOrlando Ortiz JimenezNoch keine Bewertungen
- Chapt 6Dokument15 SeitenChapt 6morphos777Noch keine Bewertungen
- Estrada - Graph Theoretic Approaches To Atomic Vibrations in FullerenesDokument43 SeitenEstrada - Graph Theoretic Approaches To Atomic Vibrations in FullerenesChern ChuangNoch keine Bewertungen
- 2008 - GDM - Physica A 387 (2008) 418-424Dokument7 Seiten2008 - GDM - Physica A 387 (2008) 418-424Vagaf1825Noch keine Bewertungen
- Decision ScienceDokument4 SeitenDecision Scienceharsha faraiyyaNoch keine Bewertungen
- Cellular Automata With Regular Behavior: Om Plex SDokument15 SeitenCellular Automata With Regular Behavior: Om Plex SCarlos ReyesNoch keine Bewertungen
- BF 01759355Dokument18 SeitenBF 01759355Ерген АйкынNoch keine Bewertungen
- Exercises On Significance ZDokument5 SeitenExercises On Significance ZMd HassanNoch keine Bewertungen
- Age DatingDokument25 SeitenAge DatingmksayshiNoch keine Bewertungen
- Quantum Optics Shot-Noise Limit of Interferometry: M. Brune, A. Aspect Homework of Lesson 6Dokument7 SeitenQuantum Optics Shot-Noise Limit of Interferometry: M. Brune, A. Aspect Homework of Lesson 6bruhbruhbruhNoch keine Bewertungen
- Horst 1982Dokument14 SeitenHorst 1982Cristian ValdezNoch keine Bewertungen
- Manual FadwaveDokument14 SeitenManual FadwaveDedi ApriadiNoch keine Bewertungen
- The New Prime Theorem 10 There Are Finite Mersenne Primes and There Are Finite Repunits PrimesDokument2 SeitenThe New Prime Theorem 10 There Are Finite Mersenne Primes and There Are Finite Repunits PrimeskgskgmNoch keine Bewertungen
- EvolutionaryStrategies BaeckDokument6 SeitenEvolutionaryStrategies BaeckSurya MahadiNoch keine Bewertungen
- Barker - Kezlan (1990) - Automorphisms of Algebras of Upper Triangular Matrices - ArchMathDokument6 SeitenBarker - Kezlan (1990) - Automorphisms of Algebras of Upper Triangular Matrices - ArchMathSandipan PaulNoch keine Bewertungen
- Axioms of ProbailityDokument5 SeitenAxioms of Probailitymonyeidavid13Noch keine Bewertungen
- GWAS ExamplesDokument4 SeitenGWAS ExamplesAbaidullahNoch keine Bewertungen
- Probability Exam UPFDokument3 SeitenProbability Exam UPFDani GonzálezNoch keine Bewertungen
- Scan Apr 16, 2022Dokument10 SeitenScan Apr 16, 2022Ashish GuptaNoch keine Bewertungen
- Hardy Weinberg Notes RevisedDokument15 SeitenHardy Weinberg Notes Revisedrichmond12proNoch keine Bewertungen
- Bounds of Operator Functions and Furuta Inequalities: Publ. RIMS, Kyoto UnivDokument8 SeitenBounds of Operator Functions and Furuta Inequalities: Publ. RIMS, Kyoto Univtdtv0204Noch keine Bewertungen
- Fundamentals of Biostatistics 8th Edition Rosner Solutions ManualDokument38 SeitenFundamentals of Biostatistics 8th Edition Rosner Solutions Manualshanerussellqe03lw100% (17)
- Chi Squared TestDokument12 SeitenChi Squared Testapi-188431847Noch keine Bewertungen
- Ion Pair FormationDokument10 SeitenIon Pair FormationrolloNoch keine Bewertungen
- For Students: Solutions To Odd Numbered End of Chapter ExercisesDokument72 SeitenFor Students: Solutions To Odd Numbered End of Chapter Exercisesxandercage100% (1)
- Independence and Bernoulli Trials (Euler, Ramanujan and Bernoulli Numbers)Dokument40 SeitenIndependence and Bernoulli Trials (Euler, Ramanujan and Bernoulli Numbers)hari_shankar55100% (1)
- Dedicated To Professor Dr. Ioan A. Rus On His 70th BirthdayDokument17 SeitenDedicated To Professor Dr. Ioan A. Rus On His 70th Birthdayfahri_amirullahNoch keine Bewertungen
- Dr. Akram Spring 2015: Basic ConceptsDokument7 SeitenDr. Akram Spring 2015: Basic Conceptsصدام حسینNoch keine Bewertungen
- Cabudol, Angel B. Bsed Ii - Math: Rules in Solving Euler Phi Function or Totient FunctionDokument4 SeitenCabudol, Angel B. Bsed Ii - Math: Rules in Solving Euler Phi Function or Totient FunctionCabudol Angel BaldivisoNoch keine Bewertungen
- Worked Examples 1Dokument13 SeitenWorked Examples 1Anonymous WyTCUDyW0% (1)
- Q4 1. LU Factorization Without PivotingDokument5 SeitenQ4 1. LU Factorization Without Pivotingsaujanya raoNoch keine Bewertungen
- 1994 Mathematica Laplace InvDokument7 Seiten1994 Mathematica Laplace InvWeny AstutiNoch keine Bewertungen
- International Centre For Theoretical PhysicsDokument6 SeitenInternational Centre For Theoretical PhysicsAlejandro Mejia RicoNoch keine Bewertungen
- Econometrics Regression Analysis With Time Series Data: Serial Correlation and HeteroskedasticityDokument34 SeitenEconometrics Regression Analysis With Time Series Data: Serial Correlation and HeteroskedasticitySurbhi GuptaNoch keine Bewertungen
- Discrete Series of GLn Over a Finite Field. (AM-81), Volume 81Von EverandDiscrete Series of GLn Over a Finite Field. (AM-81), Volume 81Noch keine Bewertungen
- Tables of Coulomb Wave Functions: Whittaker FunctionsVon EverandTables of Coulomb Wave Functions: Whittaker FunctionsNoch keine Bewertungen
- Primary Eye Examination: A Comprehensive Guide to DiagnosisVon EverandPrimary Eye Examination: A Comprehensive Guide to DiagnosisJong-Soo LeeNoch keine Bewertungen
- Radically Elementary Probability Theory. (AM-117), Volume 117Von EverandRadically Elementary Probability Theory. (AM-117), Volume 117Bewertung: 4 von 5 Sternen4/5 (2)
- 01 01Dokument14 Seiten01 01fbihansipNoch keine Bewertungen
- ComplicationsDokument3 SeitenComplicationsfbihansipNoch keine Bewertungen
- ComplicationsDokument3 SeitenComplicationsfbihansipNoch keine Bewertungen
- Clinical Manifestation EditDokument2 SeitenClinical Manifestation EditfbihansipNoch keine Bewertungen
- WHO HQ Reports G2 PROD EXT TBFinancingCountryProfileDokument1 SeiteWHO HQ Reports G2 PROD EXT TBFinancingCountryProfilefbihansipNoch keine Bewertungen
- Bls PatientDokument293 SeitenBls PatientSyarifah Alfi A. ANoch keine Bewertungen
- Monohybrid CrossDokument1 SeiteMonohybrid CrossfbihansipNoch keine Bewertungen
- Kadek AditDokument1 SeiteKadek AditfbihansipNoch keine Bewertungen
- Kadek Adit WiryadanaDokument3 SeitenKadek Adit WiryadanafbihansipNoch keine Bewertungen
- Kadek AditDokument1 SeiteKadek AditfbihansipNoch keine Bewertungen
- Exam 4Dokument10 SeitenExam 4fbihansipNoch keine Bewertungen
- 2bacterial KeyDokument3 Seiten2bacterial KeyfbihansipNoch keine Bewertungen
- 13 - Heart PhysDokument8 Seiten13 - Heart PhysfbihansipNoch keine Bewertungen