Beruflich Dokumente
Kultur Dokumente
Identification of An Unknown Amino Acid
Hochgeladen von
VanandiOriginaltitel
Copyright
Verfügbare Formate
Dieses Dokument teilen
Dokument teilen oder einbetten
Stufen Sie dieses Dokument als nützlich ein?
Sind diese Inhalte unangemessen?
Dieses Dokument meldenCopyright:
Verfügbare Formate
Identification of An Unknown Amino Acid
Hochgeladen von
VanandiCopyright:
Verfügbare Formate
BC367 Experiment 1 Identification of an Unknown Amino Acid Introduction As the building blocks of proteins, amino acids play a key
cellular role in structure and function. Proteins themselves participate in nearly every physiological event in the cell. In order to understand acid-base properties of proteins and their behavior as polyionic macromolecules, we will begin by investigating the properties of their constituent amino acids. Since all amino acids contain at least one amino and one carboxyl group, they are classified as amphoteric substances (meaning that they can act as either an acid or as a base). Such a molecule reacts with acids as follows:
R H3 N
+
K1
+
R H3 N
+
CH COO- + H3O
CH COOH + H2O
Zwitterion
Acid
Cationic form
and with bases as follows:
R H3 N
+
K2
-
R H2N CH COO-
CH COO
+ OH
+ H2O
Zwitterion
Base
Anionic form
The ionic form of the amino acid present in an aqueous solution depends on the pH of the solution. In this experiment, you will identify an unknown amino acid through acid-base titration. Titration curves of amino acids are very useful for identification. As you can see in the example for glycine shown below, a simple amino acid has two dissociation steps corresponding to loss of H+ from the acidic carboxyl group at low pH followed by loss of H+ from the more basic amino group at high pH. The pKa value for each dissociable group of an amino acid can be determined from such a titration curve by extrapolating the midpoint of each buffering region (the plateau) in the titration curve. The diagram also shows that there is a point in the curve where the amino acid behaves as a "neutral" salt. At this pH, the amino acid is predominantly a zwitterion with a net charge of zero. This point of the titration curve is the isoelectric point (pI) and can be approximated as halfway between the two points of strongest buffering capacity (the two pKa values). The isoelectric point (pI) can be estimated by: pI = 1/2 (pK1 + pK2) where K1 and K2 are the dissociation constants of the carboxyl and amino groups, respectively. pK1 of glycine is 2.35; pK2 is 9.78. Thus, the pI = 1/2 (2.35 + 9.78) = 6.06, meaning that at pH 6.06, glycine has no net charge.
BC 367, Experiment 1, Fall 2009
Theoretical titration curve of glycine with NaOH
Charged amino acids have acidic or basic side chains (R-groups) giving them more than two dissociable H+ ions. For example, glutamic acid has a carboxylic acid side chain in addition to its -carboxyl and -amino groups. A titration curve for glutamic acid will be somewhat more complex than that for glycine. Three plateau regions and three pKa values will be observed for glutamic acid: two in the acidic pH region, pK1 (-carboxyl group) = 2.2; pK2 (-carboxyl group) = 4.3; and one in the basic pH region, pK3 (-amino group)= 9.7. Members of the basic family of amino acids, such as lysine, will also exhibit three pKa values; however, due to the extra amino group they will have one pKa in the acidic pH region and two pKa values in the basic pH region. The pH at which the net charge of an amino acid is zero is called the isoelectric point, or the pI. To determine this value, average the two pKa values that flank the neutral species. Thus, the pI for glutamic acid = 1/2 (2.19 + 4.25) = 3.22. At the pI, the -carboxyl group is a negatively charged carboxylate ion, the -amino group is a positively charged ammonium ion, and the carboxyl group is a neutral protonated acid. Thus, titration curves are helpful in the identification of amino acids as follows: 1. The number of pKa values differentiates polar and nonpolar amino acids from charged amino acids. 2. The position of the pKa values for charged amino acids allows one to identify positively charged from negatively charged amino acids. 3. Comparisons between experimental and literature pKa values can allow the identification of a specific amino acid.
BC 367, Experiment 1, Fall 2009
Experimental Procedure You will perform automated titrations of both a known standard (0.1 M glycine) and an unknown amino acid (also at 0.1 M) with 1.0 M NaOH. You will use your data to determine the pKa and pI values of each amino acid, thereby allowing you to deduce the identity of your unknown (and those of your classmates). Be sure to note the code number of your unknown IN YOUR NOTEBOOK. A list of possible unknowns will be posted in the lab. Follow the instructions for the Vernier LabPro device (see the appendix) to perform your titrations. As directed, bring the pH of your amino acid solution to ~1.5 before you begin titrating, then titrate with 1.0 M NaOH. (All amino acid solutions have been brought to a pH of ~7 for this experiment.) If you think you have overlapping peaks for your unknown, repeat the titration with 0.1 M titrant. If the area of overlap is at high pH, start with the prepared amino acid solution at pH ~7 and titrate up with 0.1 M NaOH. If the area of overlap is at low pH, start with the prepared solution at pH ~7 and titrate down with 0.1 M HCl. This may help you to resolve fine detail about the positions of the overlapping peaks. (You can tell for sure whether you have two or three titratable groups after the first titration of your unknown by calculating the approximate number of equivalents needed to reach pKa1 and pKa2.) Analysis The pKa values of weak acids can generally be determined from titration curves with good accuracy. An equivalence point is often distinguished by a sharp rise in the slope of the titration curve or a peak in the derivative plot. An equivalence point is preceded by a plateau region where little change in pH is observed with the addition of acid or base. From an inspection of the Henderson-Hasselbalch equation, pH = pKa + log [A-]/[HA] it can be seen that pH= pKa at the 50% point of a titration because [conjugate base]/[acid] =1. The 50% point of titration of a particular species should correspond to the midpoint of the plateau region. An amino acid will have at least two titratable species (the -carboxyl and amino groups) and possibly three (an extra carboxyl or amino group in the side chain). Thus, two or more plateau regions should be observed, the midpoints of which correspond to the pKa values of the titratable groups. Note that when titrating a molecule that contains more than one ionizable H+, the plateau region for each ionizable H+ present will be distinct if their pKa values are more than two pH units apart (K1 > 100 K2). If the pKa values differ by less than two pH units, the titration curves will be poorly defined and may overlap to the extent that there is no point of inflection between the two curves; that is, the titration of the second species begins before the titration of the first is complete. Such an overlap of titration curves can usually be detected because of an unreasonably long plateau region that consumes two molar equivalents of titrant. Therefore, you should make sure that you calculate the equivalents of titrant added to help substantiate the identity of your amino acid. You will then have to eyeball the pKa values of overlapping plateaus. In addition to the pKa values, calculate the isoelectric point (pI) of your unknown. Compare the pKa and pI values of your unknown amino acid to literature values. On your titration plot in
BC 367, Experiment 1, Fall 2009
your notebook, label the species found on each region of the curve. Put an annotated copy of your unknown amino acid, with your name and unknown number clearly noted, on the fileserver within 24 hours after the completion of the lab period. For your write-up, include a graph for glycine and each unknown in your notebook with your assignment of its identity, points of interest labeled on the curve, and the chemical structures of the species found on each region of the curve. In your discussion thoroughly justify the identity of each unknown amino acid on the basis of the titration data. If you had a mixture of unknown amino acids, how might you identify its components?
Appendix: Instructions for the Vernier LabPro INTRODUCTION:
The Vernier LabPro is a versatile data collection interface that can be used in many different ways in the classroom and in the field. In this experiment, the Vernier LabPro is connected to a computer and used for an automated titration of your amino acids. The titration is monitored via changes in pH using a pH electrode.
INSTRUCTIONS FOR USE:
A. Make sure that the stir plate and the power supply for the DCU pump unit are plugged in and on. A blinking green light occurs when the interface has power. The Vernier system will complete series of (7) beeps as the system completes a self check. If no other beeping sounds are given by the interface, then the system is ready for use. The pump should be connected to the DIG/Sonic1 port and an electrode amplifier with an attached pH electrode should be connected to CH1. B. Calibration of the pump (finding the volume of one drop delivered by the pump): 1. Open the LoggerPro program on the dock and then open the LoggerProExperiments folder on the desktop. Double click on the icon one minute pump cal.cmbl. The pump will click and a blinking green light(s) will be visible on the interface when recognized by the software. (The toggle switches on the pump should always be set on the computer and DCU settings for the software to communicate to the pump. This should already be taken care of.) NOTE: If the equipment is not set up properly, a Sensor Confirmation window will be displayed. Be sure the sensors are securely attached to the interface and in the correct ports. If the window is not displayed, continue on. The main calibration window should be activated and the Collect button found in the top menu bar will be colored green. The table to the left will record collected data in the appropriate columns. The smaller box below the table will count the pulses delivered by the pump. The graph will display a green signal for each pulse.
BC 367, Experiment 1, Fall 2009
2. To prime the pump with deionized water, find the inlet tubing (marked as IN on the pump) and submerge the tubing into a clean 100-mL beaker of fresh dH2O. To prevent air bubbles from entering the line, make sure that the line is secure and that the opening of the tubing is at the bottom of the beaker. Put the outlet tubing into a waste beaker. 3. Click on the green Collect button. A window will appear that will ask what you would like to do: *erase and continue: will erase the last set of pulses and start at zero *cancel: cancel or exit *append to latest: will add another series of pulses to the last one *store latest run: save data in the table before starting another run NOTE: Each run is a series of 30 pulses. 4. Click on erase and continue. The pump will run for 30 pulses (refer to the small box under the table). Click on Collect again and then append to latest to add another series of 30 pulses. It will take about 90 pulses to prime the pump with water. Check for bubbles in the tubing. If you see bubbles, check your set up and add another set of pulses. If no bubbles are apparent in the lines, then continue. 5. To calibrate the pump (volume in L delivered by each pulse) find the mass of a clean dry weigh bottle (no top) or small beaker to 0.1 mg. Use a clean, dry Kim Wipe to handle the bottle because fingerprints and dirt will add to the mass and result in calibration error. Set the outlet tubing into the weigh bottle and deliver 60 pulses of deionized water to the bottle. Mass the bottle again. Using the density of water at room temperature (0.9982 g/ml at 20C), convert the mass of the water delivered to the corresponding volume of water delivered. Calculate the volume of water delivered in a single pulse by the pump (in L). 6. Remove the inlet tubing from the beaker of water. Empty the line by delivering 30-60 pulses (of air). Place the outlet tubing into a waste beaker and the inlet tubing into a 100 mL beaker of 1.0 M NaOH (or other desired titrant). Prime the pump with NaOH. Be sure that there are no bubbles in the line. 7. Exit the calibration program by using the LoggerPro menu in the main menu bar Quit LoggerPro C. To set up a titration using Vernier LabPro: 1. Open the LoggerPro program on the dock and then open the LoggerProExperiments folder on the desktop. Double click on the icon acidbasetitration.cmbl. The acidbasetitration.cmbl titration window will be displayed. As with the calibration software, the table on the left will record collected data in columns labeled with colorcoded headings. A small window under the table will display pH (live reading). Temperature will be displayed if the temperature probe is connected to the interface. The larger graphing window will display your data as it is being recorded by the interface.
BC 367, Experiment 1, Fall 2009
The X axis is the volume (L) delivered by the pump and is set from 0 to 20,000 L. The Y axis is the sensor output and is set to record pH 0 to 12. (NOTE: If necessary, the axes may be rescaled at anytime by clicking on the minimum/maximum value and typing in the preferred value.) Clear data under the Data Menu in the Logger Pro Menu bar to erase stored data and clear the window. 2. To calibrate the volume (X) axis, use the scroll bar at the bottom of the table, to find the volume column and double click on the heading. A Calculated Column window will appear. Find the equation box and change ONLY the last number in the equation to the calibrated pulse volume in L that your pump delivers. Click Done. 3. To calibrate the pH electrode chose the Experiment menu from the LoggerPro menu bar and choose Calibrate. Sliding to the right choose LabPro: 1CH1: Electrode Amplifier. The sensor settings window will appear. Make sure that the current calibration is set for Electrode Amp pH<Sensor Page 1>. Click on the calibrate now button. This is a two-point calibration. Remove the pH electrode from the electrode storage solution, make sure that the hole in the blue collar is open, rinse well with deionized water and place in the pH 4.00 (pink) buffer solution. Always make sure the ground glass frit is covered by solution and no bubbles are caught in the protective cage. Once the volts reading has stabilized, type 4.00 into the enter value box and press the Keep button. Repeat to calibrate in pH 10.00 buffer solution and press the Keep button. Press Done. 4. Add 50 mL of deionized water to a clean, dry 100-mL beaker (lines on beaker are OK). Using a graduated cylinder, add 10 mL of amino acid to the water, place the beaker on the stir plate, and add a stir bar. Rinse and dry the pH electrode and carefully place it in the beaker. Rest the electrode in the extended portion of the lip of the beaker. Carefully turn the stir plate on and arrange the beaker so that the stir bar does not hit the electrode cage. Adjust the pH to ~1.5 with 1.0 M HCl. Using the provided clip, attach the pump outlet tube to the beaker so that the end of the tubing is right at the level of the solution being tested. 5. Press the green Collect button in the main window menu bar to start the titration. For the first titration, click erase and continue. The data will be recorded in the table and the titration and derivative plots will automatically be displayed on the graph. NOTE: If this is not true, double click on the Sensor Output Y axis label and choose More. Then check the pH and dpH/dV boxes for the appropriate run. Remove the check for any unwanted data being displayed on the graph. 6. Press Stop when the pH has reached ~12-13 to end the titration (Stop is where the Collect button was at the start of the run). The color of the curves will match the color of the headings to the columns in the table. Choose File from the Logger Pro Main window menu bar and Save to assure that the data is not lost while preparing the next. 7. When all of the titration experiments are completed, and have been saved, the graph can be printed by choosing File from the LoggerPro main window menu bar and Print
BC 367, Experiment 1, Fall 2009
Graph. Entering the information in the Printing Options window will provide a label to the hardcopy of your graph, if the print footer box is checked. 8. To analyze the data points of interest on the pH and derivative plots, press the X= (also called the Examine) button in the LoggerPro titration window menu bar. The color of the data values will match the color of the plotted data. Any point along the curves can be evaluated (also recorded in the table). Record the data points that are important to your experiment. Click on the X= button again to exit data analysis. 9. You can also press the Control-Option keys on the keyboard while holding down the mouse with the cursor placed on the graph. Choosing copy from the menu box will allow the graph to be pasted into a Word document. Open Word and use Paste under the Edit menu to copy the graph. The graph can be further edited in Word. If you use the Save as option to save the data, you would need the LoggerPro Software on your computer to reopen the file. 10. To exit the program, choose Quit LoggerPro from the LoggerPro menu bar. Press Save. NOTE: When clearing data and/or erasing and continuing between titrations, always check that the pump calibration volume (pulse volume) has not been changed to the default value (it should be set the volume that you determined previously). NOTE: IF YOU ARE FINISHED WITH THE VERNIER SYSTEM, THE TITRANT MUST BE FLUSHED FROM THE PUMP: Reopen the one minute pump calibration.cmbl and place the inlet tubing into a beaker of deionized water. Place the outlet tubing in a waste beaker. Be sure that the line is secure and that the open end of the tubing is placed at the bottom of the beaker to reduce bubbles in the line. Use at least 90 pulses to run deionized water through the pump and tubing. Then place the inlet hose on the bench top and use 60 pulses to dry the lines. Please rinse the pH electrode and place it in the electrode storage solution (the hole in the blue collar should be open). Unplug the system (both plugs) when done.
Das könnte Ihnen auch gefallen
- Handbook of Coordination Catalysis in Organic ChemistryVon EverandHandbook of Coordination Catalysis in Organic ChemistryNoch keine Bewertungen
- Experiment 2 - Adsorption of Liquids Onto Solid Surfaces: TheoryDokument3 SeitenExperiment 2 - Adsorption of Liquids Onto Solid Surfaces: TheoryfrankjenNoch keine Bewertungen
- Experiment 4 FWRDokument5 SeitenExperiment 4 FWRSarah HermosuraNoch keine Bewertungen
- Lab ManualDokument19 SeitenLab Manualanon_467104036Noch keine Bewertungen
- Experiment 2 Uv-Visible Determination of An Unknown Concentration of Kmno Solution Theory/BackgroundDokument13 SeitenExperiment 2 Uv-Visible Determination of An Unknown Concentration of Kmno Solution Theory/BackgroundMuhammad Azri HaziqNoch keine Bewertungen
- Lab Ins 1 (Spectronic 20)Dokument7 SeitenLab Ins 1 (Spectronic 20)Ayale Mn100% (3)
- Anal Chem 3 - Test 1-2016Dokument4 SeitenAnal Chem 3 - Test 1-2016Buhle BuhleNoch keine Bewertungen
- 11 Fruit JuicesDokument8 Seiten11 Fruit JuicesthangesspNoch keine Bewertungen
- Measurement of The Gas Constant and Molar Volume of Oxygen GasDokument12 SeitenMeasurement of The Gas Constant and Molar Volume of Oxygen GasJennifer Im0% (1)
- EXPERIMENT 3a and 3b - Aluminum Content Via Redox and ColorimeterDokument13 SeitenEXPERIMENT 3a and 3b - Aluminum Content Via Redox and ColorimeterTrupti soniNoch keine Bewertungen
- Lab Report 10 Organic Chemistry UVA 2411Dokument6 SeitenLab Report 10 Organic Chemistry UVA 2411Alia LieNoch keine Bewertungen
- Dms 111 Manual by Michael K. Chirchir and Githii WainainaDokument173 SeitenDms 111 Manual by Michael K. Chirchir and Githii WainainaAdventist NaturopathyNoch keine Bewertungen
- Exp 1Dokument11 SeitenExp 1Zhyhui OngNoch keine Bewertungen
- 15 - Aldehyde and KetonesDokument66 Seiten15 - Aldehyde and KetonesIrfan Raza100% (1)
- AP Chemistry - Acid Dissociation Constant Ka LabDokument4 SeitenAP Chemistry - Acid Dissociation Constant Ka LabJonathan Chen83% (6)
- Preparation of Tetraamminecopper II Sulphate.Dokument10 SeitenPreparation of Tetraamminecopper II Sulphate.DaizLee Ahmad25% (4)
- Experiment No 18Dokument4 SeitenExperiment No 18Suvrasoumya Mohanty100% (2)
- Preparation of 4-Methylcyclohexene From Dehydration of 4-MethylcyclohexanolDokument8 SeitenPreparation of 4-Methylcyclohexene From Dehydration of 4-MethylcyclohexanolHidayu AdnanNoch keine Bewertungen
- CHE555 Assignment 1 Mac 2015Dokument2 SeitenCHE555 Assignment 1 Mac 2015Jaja TeukieNoch keine Bewertungen
- Revised Jobs MethodDokument5 SeitenRevised Jobs Methodsilwadi71Noch keine Bewertungen
- Titration Phosphoric AcidDokument1 SeiteTitration Phosphoric AcidKiany SirleyNoch keine Bewertungen
- Kinetics of Ester Hydrolysis NewDokument3 SeitenKinetics of Ester Hydrolysis Newbits_who_am_iNoch keine Bewertungen
- Vibration - Rotation Spectroscopy of HCL and DCLDokument9 SeitenVibration - Rotation Spectroscopy of HCL and DCLAngela LamasNoch keine Bewertungen
- Determination of Vitamin CDokument2 SeitenDetermination of Vitamin CWalwin HareNoch keine Bewertungen
- Unit 1. Chemoselectivity and Protecting GroupsDokument84 SeitenUnit 1. Chemoselectivity and Protecting GroupsBenjamín BohiguesNoch keine Bewertungen
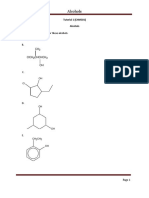
- Tutorial 1 SolutionsDokument20 SeitenTutorial 1 Solutionsanushka shagunNoch keine Bewertungen
- CHM 260 Laboratory Report: Experiment 2: Uv Visible Determination of An Unknown Concentration of Kmno4 SolutionDokument11 SeitenCHM 260 Laboratory Report: Experiment 2: Uv Visible Determination of An Unknown Concentration of Kmno4 SolutionAwathif Wawa100% (1)
- Phys Chem 3 - ElectrochemistryDokument26 SeitenPhys Chem 3 - ElectrochemistryClement ThabangNoch keine Bewertungen
- NMR Kinetics: Study of A Reversible Hydrolysis ReactionDokument8 SeitenNMR Kinetics: Study of A Reversible Hydrolysis ReactionOldbooklover100% (2)
- Experiment 5 AASDokument15 SeitenExperiment 5 AASnn bbNoch keine Bewertungen
- Equilibrium Lab ReportDokument3 SeitenEquilibrium Lab ReportJustin G-Hood Jung100% (2)
- Acid Dissociation ConstantDokument4 SeitenAcid Dissociation ConstantJair RangelNoch keine Bewertungen
- Chemistry of Carbonyl CompoundsDokument28 SeitenChemistry of Carbonyl CompoundsRhondene WintNoch keine Bewertungen
- Experiment Baking SsodaDokument7 SeitenExperiment Baking Ssodaatynzaty0% (1)
- Voltaic Cell Lab ReportDokument5 SeitenVoltaic Cell Lab Reporthelia bazarganNoch keine Bewertungen
- Practical Analytical 1 ,,chemistryDokument45 SeitenPractical Analytical 1 ,,chemistryFadlin AdimNoch keine Bewertungen
- Parallel Plate Capacitor Lab ReportDokument5 SeitenParallel Plate Capacitor Lab ReportSyamil Amir HamzahNoch keine Bewertungen
- EXPERIMENT 2 Reduction of CamphorDokument2 SeitenEXPERIMENT 2 Reduction of CamphorDania FaridNoch keine Bewertungen
- 3 Determination of Complex Ion by Jobs MethodDokument2 Seiten3 Determination of Complex Ion by Jobs Methodvishwanathz47Noch keine Bewertungen
- Uv Vis Spectroscopy Lab Report of The Detection of Analytes From Environmental Samples Using UvDokument13 SeitenUv Vis Spectroscopy Lab Report of The Detection of Analytes From Environmental Samples Using UvSaba Naseer100% (1)
- Quantitative Determination of Phosphorus in Plant Food Using Household ChemicalsDokument3 SeitenQuantitative Determination of Phosphorus in Plant Food Using Household ChemicalsMaryNoch keine Bewertungen
- Hydrolysis of Tert-Butyl Chloride and Solvent EffectDokument7 SeitenHydrolysis of Tert-Butyl Chloride and Solvent EffectangelbenavidezNoch keine Bewertungen
- Cape Chemistry Lab CompressDokument6 SeitenCape Chemistry Lab CompressDesmond JonesNoch keine Bewertungen
- 2D NMRDokument10 Seiten2D NMRHariprasad Reddy100% (1)
- Preparation and Reaction of Carboxylic AcidsDokument6 SeitenPreparation and Reaction of Carboxylic AcidsIndhumathiNoch keine Bewertungen
- Equilibrium Constant For Hydrolysis Lab6finalDokument9 SeitenEquilibrium Constant For Hydrolysis Lab6finalapi-534386927Noch keine Bewertungen
- Punjab College Pattoki: Spring 2021: Course Outline Bs Program Semester 6ThDokument8 SeitenPunjab College Pattoki: Spring 2021: Course Outline Bs Program Semester 6ThFareeha ShakeelNoch keine Bewertungen
- Laboratory Manual: SKT 1013 Introduction To Inorganic ChemistryDokument23 SeitenLaboratory Manual: SKT 1013 Introduction To Inorganic Chemistryizz isalahNoch keine Bewertungen
- Indus Waste ProblemsDokument3 SeitenIndus Waste ProblemsZeus Ian DuarteNoch keine Bewertungen
- ST01502 Earth Science (Sains Bumi)Dokument22 SeitenST01502 Earth Science (Sains Bumi)Moganambal RavichandranNoch keine Bewertungen
- Tutorial 1 - Alcohol PDFDokument5 SeitenTutorial 1 - Alcohol PDFNurul Athirah JainiNoch keine Bewertungen
- Gas ChromatographyDokument19 SeitenGas ChromatographyNauman Mithani100% (2)
- Analytical Chemistry Notes IiDokument9 SeitenAnalytical Chemistry Notes IiJabez MatigaNoch keine Bewertungen
- Discussions Exp 14 RecrystallizationDokument4 SeitenDiscussions Exp 14 RecrystallizationEdwin fooNoch keine Bewertungen
- 14 - Lab 14 - R-HPLC For Detn of CaffeineDokument7 Seiten14 - Lab 14 - R-HPLC For Detn of CaffeineHoang Huong TraNoch keine Bewertungen
- Enzymes: A Practical Introduction to Structure, Mechanism, and Data AnalysisVon EverandEnzymes: A Practical Introduction to Structure, Mechanism, and Data AnalysisBewertung: 4 von 5 Sternen4/5 (2)
- Coordination Chemistry: Invited Lectures Presented at the 20th International Conference on Coordination Chemistry, Calcutta, India, 10-14 December 1979Von EverandCoordination Chemistry: Invited Lectures Presented at the 20th International Conference on Coordination Chemistry, Calcutta, India, 10-14 December 1979D. BanerjeaNoch keine Bewertungen
- 5 Useful Tables For Bacterial IdentificationDokument4 Seiten5 Useful Tables For Bacterial IdentificationVanandiNoch keine Bewertungen
- Extraction of Genomic DNA From Yeasts For PCR-based ApplicationsDokument3 SeitenExtraction of Genomic DNA From Yeasts For PCR-based ApplicationsScientificBoyNoch keine Bewertungen
- Coagulase TestDokument3 SeitenCoagulase TestVanandiNoch keine Bewertungen
- Chem 240 Lab Manual - 2013Dokument56 SeitenChem 240 Lab Manual - 2013VanandiNoch keine Bewertungen
- Experiment 2Dokument5 SeitenExperiment 2VanandiNoch keine Bewertungen
- Wino Gradsky 04 ExperimentDokument4 SeitenWino Gradsky 04 ExperimentVanandiNoch keine Bewertungen
- PH CalculationsDokument4 SeitenPH CalculationsVanandiNoch keine Bewertungen
- Molecule Structure Point Group Symmetry Elements Symmetry OperationsDokument6 SeitenMolecule Structure Point Group Symmetry Elements Symmetry OperationsVanandiNoch keine Bewertungen
- Practice ExamDokument9 SeitenPractice ExamVanandi100% (1)
- Golden Sun The Lost Age Walkthrough From GamefaqsDokument104 SeitenGolden Sun The Lost Age Walkthrough From GamefaqsVanandiNoch keine Bewertungen
- Web Lecture CHP 20 Section 1Dokument9 SeitenWeb Lecture CHP 20 Section 1VanandiNoch keine Bewertungen
- CHM 218 - Introduction To Inorganic Chemistry Spring 2003 IpfwDokument18 SeitenCHM 218 - Introduction To Inorganic Chemistry Spring 2003 IpfwVanandiNoch keine Bewertungen
- HSABDokument1 SeiteHSABVanandiNoch keine Bewertungen
- UT Dallas Syllabus For Chem2401.001 05f Taught by Paul Pantano (Pantano)Dokument5 SeitenUT Dallas Syllabus For Chem2401.001 05f Taught by Paul Pantano (Pantano)UT Dallas Provost's Technology GroupNoch keine Bewertungen
- Free Energies of Proton-Coupled Electron Transfer Reagents and Their ApplicationsDokument49 SeitenFree Energies of Proton-Coupled Electron Transfer Reagents and Their ApplicationsZongxin JinNoch keine Bewertungen
- JEE CHemistry FormulasDokument18 SeitenJEE CHemistry FormulasVishant Kamble100% (1)
- Anaerobic Digestion of Tuna Waste For The Production of Volatile Fatty AcidsDokument7 SeitenAnaerobic Digestion of Tuna Waste For The Production of Volatile Fatty AcidsLAURA DANIELA CARDONA ACUNANoch keine Bewertungen
- 222-10 Exam #1 Answers PDFDokument8 Seiten222-10 Exam #1 Answers PDFbrich22Noch keine Bewertungen
- Script For The Reporting in ChemDokument11 SeitenScript For The Reporting in ChemJamaica SalvadorNoch keine Bewertungen
- Lysotracker and Lysosensor ProbesDokument4 SeitenLysotracker and Lysosensor Probesplastioid4079Noch keine Bewertungen
- Essential Organic Chemistry Canadian 3rd Edition Bruice Test BankDokument14 SeitenEssential Organic Chemistry Canadian 3rd Edition Bruice Test Bankkatherinelynchiozjftcaqw100% (9)
- Experiment 6 ExtractionDokument10 SeitenExperiment 6 Extractionwallace120Noch keine Bewertungen
- Solubilizing Excipients in Oral and Injectable Formulations-REVIEW-VERY IMPORTANTDokument30 SeitenSolubilizing Excipients in Oral and Injectable Formulations-REVIEW-VERY IMPORTANTraju1559405Noch keine Bewertungen
- CH 2 - ProblemsDokument6 SeitenCH 2 - ProblemsKhris Griffis94% (17)
- Acid-Base Indicators For Use in TitrationDokument7 SeitenAcid-Base Indicators For Use in TitrationMaxNoch keine Bewertungen
- Identifying of Unknown Monoprotic AcidDokument21 SeitenIdentifying of Unknown Monoprotic AcidjuaxxoNoch keine Bewertungen
- Weak Acid and Base EditedDokument52 SeitenWeak Acid and Base EditedoofNoch keine Bewertungen
- TebuconazoleDokument195 SeitenTebuconazoleKen EspinoNoch keine Bewertungen
- 7.0 Ionic EquilibriaDokument124 Seiten7.0 Ionic EquilibriaTasya KassimNoch keine Bewertungen
- TEST-17: JEE MainDokument20 SeitenTEST-17: JEE MainRdNoch keine Bewertungen
- Snap Revise NotesDokument8 SeitenSnap Revise Notestassu100% (1)
- Elsevier Pka DeterminationDokument12 SeitenElsevier Pka DeterminationJack SmitNoch keine Bewertungen
- 3 Acid Dissociation Constant of Methyl RedDokument2 Seiten3 Acid Dissociation Constant of Methyl RedErcille Mae Oraiz PacamoNoch keine Bewertungen
- Thermodynamics of Solutions and Electrochemistry: 22306 Physical Chemistry IDokument127 SeitenThermodynamics of Solutions and Electrochemistry: 22306 Physical Chemistry ISonalNoch keine Bewertungen
- The Organic Chemistry of Enzyme-Catalysed ReactionsDokument63 SeitenThe Organic Chemistry of Enzyme-Catalysed ReactionskunwarskNoch keine Bewertungen
- Preformulation and Product Development: Bishnu Prasad KoiralaDokument59 SeitenPreformulation and Product Development: Bishnu Prasad Koiralasaloni patelNoch keine Bewertungen
- OCLUE Organic Chemistry Life The Universe Amp Everything 1614802633Dokument276 SeitenOCLUE Organic Chemistry Life The Universe Amp Everything 1614802633Alex SommersNoch keine Bewertungen
- Chemistry Grade 12 New TextbookDokument248 SeitenChemistry Grade 12 New Textbookdereje dawitNoch keine Bewertungen
- Solutions To MCAT TPR Practice Test 1 23690Dokument83 SeitenSolutions To MCAT TPR Practice Test 1 23690Tony Adeosun75% (8)
- Article 7Dokument37 SeitenArticle 7Mariel :vNoch keine Bewertungen
- Module 5&6 Chemistry Notes (Created by Etho - X - BOS)Dokument25 SeitenModule 5&6 Chemistry Notes (Created by Etho - X - BOS)noorNoch keine Bewertungen
- Rate Constants of Reactions of Ozone With Organic and Inorganic Compounds in WaterDokument10 SeitenRate Constants of Reactions of Ozone With Organic and Inorganic Compounds in WaterIngrid Rincón ValdiviesoNoch keine Bewertungen
- Biochemistry Practice Exam Question and AnswersDokument5 SeitenBiochemistry Practice Exam Question and Answerspunith gowdaNoch keine Bewertungen