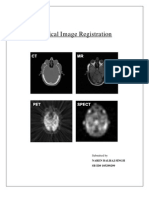
Beruflich Dokumente
Kultur Dokumente
A Visual Approach
Hochgeladen von
Rasool ReddyOriginaltitel
Copyright
Verfügbare Formate
Dieses Dokument teilen
Dokument teilen oder einbetten
Stufen Sie dieses Dokument als nützlich ein?
Sind diese Inhalte unangemessen?
Dieses Dokument meldenCopyright:
Verfügbare Formate
A Visual Approach
Hochgeladen von
Rasool ReddyCopyright:
Verfügbare Formate
©TECHPOOL STUDIOS, INC.
By Felix Ritter, Tobias Boskamp, André Homeyer, Hendrik Laue,
Michael Schwier, Florian Link, and Heinz-Otto Peitgen
S
ince the discovery of the X-ray radiation by Wilhelm (US), offers previously unattained opportunities for diagno-
Conrad Roentgen in 1895, the field of medical imaging sis, therapy planning, and therapy assessment. Medical image
has developed into a huge scientific processing is essential to leverage this increas-
discipline. The analysis of patient data ing amount of data and to explore and present
acquired by current image modali- A Visual Approach the contained information in a way suitable
ties, such as computerized tomogra- for the specific medical task.
phy (CT), magnetic resonance tomography In this tutorial, we will approach the anal-
(MRT), positron emission tomography (PET), or ultrasound ysis and visualization of medical image data in an explorative
manner. In particular, we will visually construct the image-
Digital Object Identifier 10.1109/MPUL.2011.942929
processing algorithms using the popular graphical data-flow
Date of publication: 30 November 2011 builder MeVisLab, which is available as a free download for
60 IEEE PULSE ▼ NOVEMBER/DECEMBER 2011 2154-2287/11/$26.00©2011 IEEE
noncommercial research [1]. We felt that it could be more inter- Such a separation is the most basic form of segmentation called
esting for the reader to see and explore examples of medical im- thresholding. Often, however, materials do not differ enough in
age processing that go beyond simple image enhancements. The radio density to be distinguishable by using thresholding alone.
part of exploration, to inspect medical image data and experi- For instance, the HU value of blood is around +40 HU and the
ment with image-processing pipelines, requires software that value of liver tissue is between +40 and +60 HU. The perception
encourages this kind of visual exploration. of blood-filled vascular structures within the liver would be dif-
ficult, if not impossible, given such a small difference. This is why
Image Processing contrast agents are important. When applied to a patient, they
In contrast to general image processing that has primary goals, will change the imaging characteristics of the aimed structures.
such as enhancing the esthetics of an image or creating art, the Using a contrast agent for vascular structures will increase the
sole purpose of medical image processing is to improve the in- HU value of blood significantly above the value of the surround-
terpretability of the depicted contents. This may involve an ing tissue (e.g., +120 HU). The contrast-enhanced blood will not
enhancement of the image itself to increase the only flow through the arteries and eventually ar-
perception of certain features as well as the auto- rive in the liver but also accumulate in the liver
mated or manual extraction of information. The sole purpose tissue itself. Hence, timing becomes very crucial
Since we can only discuss a few image analy- of medical image for imaging certain structures.
sis and visualization methods here, the following In contrast to the Hounsfield scale of CT im-
classification of the most important categories
processing is aging, the MRT imaging scale has no absolute
will provide a short reference for further inves- to improve the physical meaning. But like a black and a white
tigation: interpretability of the photograph in which you can easily distinguish
▼ Image enhancement: The removal of image depicted contents. a darker lawn in front of a house painted in a
distortions, such as noise and background brighter color without knowing the exact bright-
inhomogeneities, as well as the enhance- ness of each color, MRT images will depict dif-
ment of image contours and other relevant properties. ferent tissue characteristics in a meaningful way. In most cases,
▼ Image segmentation: The identification of the contours of an relative differences in image values are sufficient for medical
anatomical structure, such as an organ, a vessel, or a tumor diagnosis and absolute values are not required. Protocols for ac-
lesion. quiring absolute values often need multiple scans and are there-
▼ Image registration: The spatial transformation of one image fore also more time consuming.
such that it directly matches a given reference image. This Generally, medical images contain noise. Noise may impede
is necessary, for example, in the combined visualization of the analysis of regions based on voxel values. Smoothing the
images from different modalities (e.g., PET/CT). image may help in this case but will also reduce the perception
▼ Quantification: The determination of geometrical proper- and detection of contours. Like with other basic image enhance-
ties of an anatomical structure (e.g., volume, diameter, and ment methods such as sharpening, smoothing can be based on
curvature) or physiological properties such as perfusion a fundamental image operation called convolution. Convolution
characteristics or tissue composition. replaces each voxel value by a weighted sum of its neighbors.
▼ Visualization: Two-dimensional (2-D) and three-dimen- The weights are specified by the convolution kernel, which can
sional (3-D) rendering of image data (e.g., volume render- be represented by a matrix. The most basic smoothing kernel re-
ing) and virtual models (e.g., surface models) of organs and places each voxel with the weighted average of the direct neigh-
other anatomical structures. bors, hence will contain a value of 1 in all elements of a 3 3 3
▼ Computer-aided detection: The detection and characterization 3 3 matrix. More common, however, is a binomial (Gaussian-
of pathological structures and lesions, such as tumor lesions like) convolution kernel that applies more weight to less-distant
or vessel obstructions. neighbors.
The term image is used here not only to refer to a conven- A very common segmentation method for coherent regions of
tional, 2-D image, for example, a radiograph. An image may also an image is region growing. Region growing combines homog-
be comprised of a 3-D volume of parallel slices (e.g., a series of enous, neighboring voxels to form a region starting with a given,
CT or MRT tomograms) and even a sequence of image volumes initial voxel. Accepted values of adjacent voxels may be fixed and
acquired over time, such as a dynamic MRT study, may be con- set in the beginning or are being adapted automatically while the
sidered a four-dimensional image. algorithm spreads in the vicinity. The initial set or range of ac-
Images of volumetric imaging modalities, such as CT and cepted voxels is often derived from the value of the initial voxel.
MRT, are the most common source for medical image processing. While looking at individual slices of a volumetric image is
Each volume is composed of volumetric image elements called the most common form of diagnostic reading or inspection in
voxels that represent a value on a regular, 3-D grid. The image the preparation of a surgical intervention, 3-D visualizations
values of CT images are of absolute value measured on the Houn- may assist the analysis of spatial relations of different structures
sfield scale, a scale describing radio density. In our second ex- (Figure 1). Most 3-D visualizations are 2-D projections of a 3-D
ample, we use knowledge about the radiation absorption of dif- image displayed on a conventional screen. By adding the abil-
ferent materials to separate tissue, which has a value of around ity to interactively change the point of view and reveal certain
+30 Hounsfield units (HU) from air, approximately −1,000 HU. structures while hiding others, 3-D visualizations become an
NOVEMBER/DECEMBER 2011 ▼ IEEE PULSE 61
FIGURE 1 A coherent 2-D and 3-D visualizations of multimodal brain data using MeVisLab. A lesion (blue) as well as an arteriovenous
malformation (red) are highlighted. The crosshair indicates the same location in both visualizations.
extremely useful tool to build up spatial understanding. There an internal development tool of the German Fraunhofer MEVIS
are two main 3-D visualization methods for medical image data: research institute and its commercial spin-off MeVis Medical So-
direct volume rendering and polygonal isosurface rendering. lutions, MeVisLab was made publicly available in 2004 and is
Direct volume rendering, as the name implies, is currently used in numerous research and devel-
able to directly visualize image data. We are not opment sites worldwide.
required to segment structures beforehand since MeVisLab includes a comprehensive, extensi-
MeVisLab allows
the whole image data are evaluated for display. ble library of functional components, hereinafter
Voxel values are mapped to colors and opacity the user to create referred to as modules, for segmentation, registra-
values according to a transfer function. In con- macro modules tion, and quantitative morphological as well as
trast, polygonal isosurface rendering relies on the that encapsulate functional analysis. For the visual presentation of
ability to extract regions from the image, which a network with medical data, interactive 2-D and 3-D viewing,
are then rendered as polygonal meshes. An iso- all its modules, contour, surface model, and volume represen-
surface represents regions of voxels of a certain tations are available (Figure 1). These modules
connections, and
value. While isosurfaces may be extracted di- can be combined to form module networks that
rectly from the image, this method is commonly parameterizations. implement complex image processing and visu-
applied to mask images of already segmented alization tasks. MeVisLab uses a graphical pro-
structures. gramming approach for the development of such
The examples discussed hereinafter will combine a number networks, allowing the developer to instantiate modules in a
of image analysis and visualization methods to provide solutions network and draw connections in-between to define the flow of
to common medical tasks. data and control information (Figure 2). Besides the MeVisLab-
native MeVis image-processing library (ML), the well-known
MeVisLab National Library of Medicine Insight Segmentation and Regis-
The design and exploration of new methods in medical image tration Toolkit (ITK) as well as the Visualization ToolKit (VTK)
processing and visualization are creative processes starting with have been integrated in a manner that encourages and allows
an initial idea in need of realization and validation. In the same combination of their modules.
manner as architects sketch their ideas of a new building on a In analogy to the concept of classes in object-oriented pro-
sheet of paper, making refinements on their way to the final gramming, MeVisLab allows the user to create macro modules
drawing, the image-processing researcher could benefit from that encapsulate a network with all its modules, connections,
sketching algorithmic ideas in a visual way if this sketch can be and parameterizations. Such macro modules may be instanti-
brought to life with real medical image data. ated and combined with other modules in a network, allowing
MeVisLab provides such an environment of advanced medi- one to implement even more complex networks in a structured,
cal image processing and visualization building blocks that can hierarchical, and modular way.
be combined visually to create new algorithms and prototypi- The parameters offered by many modules to control their
cal applications, enabling the researcher or clinician to evalu- internal operations can be manipulated in each module’s
ate these algorithms in clinical settings. Originally designed as graphical user-interface (GUI) panel (Figure 2). Connections
62 IEEE PULSE ▼ NOVEMBER/DECEMBER 2011
FIGURE 2 The MeVisLab visual development environment. Algorithmic building blocks displayed as boxes can be combined to form
a more complex algorithm via an intuitive GUI. The parameter panel of the DtfSkeletonization algorithm is displayed to enable its
configuration.
between parameters can be established to automatically trans- eases, in particular, the detection of abnormal variations such
fer information between algorithms. This panel may also display as aneurysms. In surgery, estimating the risk of injuring the
result values that are generated by the module. blood supply and drainage of organs is of great
The layout and appearance of the module panels importance. The second example segments,
is defined using an abstract, hierarchical scripting quantifies, and visualizes a round lesion that
MRT images will
language. Such user-interface panels may also be is attached to a healthy tissue. Here, we would
defined for a complete network, giving access to
depict different tissue like to calculate the maximum diameter of a
only those parameters and results that are neces- characteristics in a metastasis in the human lung that connects
sary for using the functionality implemented in meaningful way. to the chest wall but does not still infiltrate it.
the network. The automatic calculation of the maximum di-
MeVisLab offers specific modules to inte- ameter in 3-D provides a much more reliable
grate a module network into the clinical data infrastructure. indication of shrinkage or growth in follow-up examinations
The well-accepted Digital Imaging and Communications in than the manual measurement of a diameter in just one cross-
Medicine (DICOM) standard is used to receive image data sectional image.
from the imaging modalities or to transfer images from or to
a clinical image data archive [picture archiving and commu- Segmentation and Visualization
nication system (PACS)]. DICOM tools, such as the popular of Vascular Structures
OsiriX DICOM viewer, may serve as an interface for MeVis- If you like to follow the discussion of the reconstruction of vas-
Lab-based image analysis. As a result, complete software ap- cular structures in the MeVisLab application, please open the
plication prototypes can be created to facilitate the evaluation network file Vascular Segmentation.mlab in [6]. In
of image analysis or visualization method as well as dedicated the left viewing panel of the example network, click with your
workflows in a clinical setting providing valuable input for the mouse in one of the bright white circular spots that represent
further development of a method. a vascular structure. Click again in another white spot to add
other vascular structures (Figure 2). To remove the markers
Image Processing Using MeVisLab you just placed, double-click on the module MarkerEditor
Two representative examples will guide our discussion on the in the network window and activate the button Delete all.
processing of medical image data using MeVisLab. Both ex- The segmented structures are indicated in an orange color in
ample networks and image data are available for download the 2-D and 3-D views.
[6] and can be explored with an installation of MeVisLab [1]. The reconstruction of vascular structures from a volumetric
The first example illustrates the segmentation of elongated data set (stack of images) involves several steps (see Figures 3
structures, such as vascular systems, and their visualization and 4):
in 3-D. The reconstruction of vascular structures from image 1) Preprocessing: Gaussian smoothing of noisy image data.
data is highly useful for the diagnosis of cardiovascular dis- 2) Segmentation: Region growing using an initial seed point.
NOVEMBER/DECEMBER 2011 ▼ IEEE PULSE 63
Preprocessing Segmentation Postprocessing Visualization Measurement
Gaussian Smoothing Region Growing Graph Analysis Volume Rendering
Diameter
Median Filtering Watersheds Surfaces
Stenoses
Vesselness Filtering Fuzzy Connectedness MPRs
Wall Thickness
Thresholding Level Sets
Dependent Tissue
Shortest Path
Tracking
FIGURE 3 An image-processing pipeline for the visualization and quantification of vascular structures from image data.
3) Postprocessing: Analysis of the segmentation object and con- data have been compacted and stored as a single file, compressed
struction of a model-based representation. 3-D data set in the MeVisLab native format. These data are acces-
4) Visualization: Surface rendering of the model-based repre- sible via the ImageLoad module. We use a variant of this mod-
sentation. ule called LocalImage to automatically locate the data in the
5) Measurement: Quantification of dependent tissue for risk es- directory of our example network. This module is a good choice
timation in surgery. for demonstration networks.
For our first example, we omit the preprocessing step and start In MeVisLab, we distinguish between image-processing
with the segmentation of the loaded image data. Loading image modules and modules that perform visualization tasks because
data can be done in several ways, depending on the nature of the of the different underlying frameworks. Whereas image-pro-
data, but like the rest of the image-processing pipeline always cessing modules (colored in blue) form a pipeline, for which
involves a module. A series of DICOM images, which is the most dedicated input and output connectors are provided, visualiza-
basic form for medical images, can be retrieved from a disk us- tion modules (colored in green) are connected in a hierarchical
ing the DirectDicomImport module. The provided example graph called a scene graph. This scene graph is traversed depth
3DView
SoCustomExaminerViewer
SoShadowCast Visualization
SoBackground SoOrientationInset
2DView
SoVascularSystem SoOrientationModel
View2D
SoView2DOverlay DtfSkeletonization Postprocessing
RegionGrowing
Segmentation
LocalImage MarkerEditor
SoView2DMarkerEditor
FIGURE 4 Segmentation, postprocessing, and visualization parts of the vascular reconstruction network.
64 IEEE PULSE ▼ NOVEMBER/DECEMBER 2011
first from left to right. At the root of this scene graph, we often an exact centerline and the radius at each voxel of the skeleton
find a viewing component displaying the visual result of the are computed. The skeletonization uses the binary segmenta-
image-processing network. tion image and successively erodes border voxels until a cen-
Visualization and analysis often involve a separation of ter voxel is found. A subsequent graph analysis transforms the
structures. We apply a 3-D region growing ap- skeleton in a directed, acyclic graph where nodes
proach to the original image data to segment represent branches (Figure 3). Further enhance-
the vascular system. Starting with an ini- MeVisLab provides UI ments may involve pruning and smoothing of
tial voxel from the contrast-enhanced image, the graph. This model has been implemented in
building capabilities
neighboring voxels of similar value are com- a MeVisLab module called DtfSkeletoniza-
bined. A 2-D slice-based visualization, com- to hide the network tion. It is directly connected to the SoVascu-
monly found in medical workstations, enables complexity from larSystem module for the final graph-based
the diagnostic exploration of the image set and the user. visualization. The module, among other things,
allows the user to place a marker on an initial allows the user to limit the rendering of the vas-
voxel using the mouse or trackpad. The seg- cular structures to a certain branching level. For
mented structures are highlighted on each slice to indicate instance, this is helpful when supporting planning of surgical
the result of the region growing. Two main parameters influ- interventions where branching levels indicate the volume of the
ence the result, the lower and upper thresholds of voxel val- dependent tissue.
ues and the initial voxel locations. Multiple markers may be The example previously shown in Figure 2 adds a shadow
placed to segment disconnected regions. The implementation and a 3-D depiction of the skull to the 3-D view to facilitate ori-
tries to minimize computation by adding to or removing from entation for nonexperts. Buttons for the three main anatomical
already segmented regions if parameters fit. planes, axial (aka transverse), sagittal, and coronal, are placed
The 3-D visualization of the segmented vascular structures at the right side of the 3-D view. For an occasional analysis of
supports the recognition of branching patterns as well as the in- image data, working directly with the MeVisLab network and
spection and analysis of their path. As outlined before, we may individual module panels might be acceptable. However, for fre-
apply direct volume rendering and polygonal isosurface render- quent use or studies run by a different person, a more usable
ing. With both methods, however, small vessels tend to disap- and comprehensible user interface (UI) will be less error prone
pear completely. Owing to the discrete nature of the images and and also more productive. MeVisLab provides UI building ca-
the segmentation mask, a model-based reconstruction algorithm pabilities to hide the network complexity from the user. Whole
that respects the thin structures of vessel trees whose diameters module panels or parts thereof can be combined with viewing
often grow and shrink from slice to slice will result in a much panels and other supporting UI components to form a custom-
smoother visualization. tailored image-processing application. Figure 5 shows the final
A simple but adequate model assumes that the cross sec- UI for this example network.
tion of nonpathologic vessels has a circular shape. Based on the The UI script is written in a simple UI language (see Figure 6)
Skeletonization algorithm with topology-preserving thinning, that references module panels or parameter fields. For instance,
FIGURE 5 The final UI of the vascular segmentation example. The complexity of the underlying module network is hidden from the
user. Choose Scripting S Start Network Script in MeVisLab to display this UI.
NOVEMBER/DECEMBER 2011 ▼ IEEE PULSE 65
the Delete All Markers but-
ton in our final UI is created 1 Window {
by referring to the param- 2 style = _default
eter field deleteAll of the 3
module MarkerEditor. 4 title = “Vessel Segmentation”
MarkerEditor is the 5
instance name of the So- 6 Horizontal {
View2DMarkerEditor 7 expandY = yes
module. The viewing panels 8 Box “Input” { expandY = yes layout = Vertical
are also referred to by the in- 9 Horizontal {
stance name but are suffixed 10
HyperLabel { text = “Scroll to a slice in which the vascular structures
by the special identifier .self. 11 you would like to segment are visible and click into
12
Layout components, such as the center of one or more vascular structures.” }
13
boxes as well as horizontal Button MarkerEditor.deleteAll { title = “Delete All Markers” }
14 }
or vertical alignment groups, 15 Viewer 2DView.self { mw = 400 mh = 400 type = SoRenderArea }
help to arrange and semanti- 16 }
cally combine interface com- 17 Box “Output” { expandY = yes layout = Vertical
ponents. To create or open 18 Viewer 3DView.self { mw = 400 }
the UI script of a MeVisLab 19 }
network, select Scripting S 20 }
Edit Network Script from 21 }
the menu bar. The script will
open in the MeVisLab text
FIGURE 6 The UI script defines which parameters of the image-processing network are visible and
editor, which provides syn- how they are presented to the user.
tax highlighting, context-
sensitive help, and automatic
completion for the modules and fields of the For an inspection of intermediate results in the
image-processing network. The result of the UI An isosurface image-processing pipeline, display the Output
script can be tested at any time within MeVis- represents regions Inspector by choosing View S Views S Out-
Lab by selecting Scripting S Start Network of voxels of a certain put Inspector from the menu bar. The Output
Script from the menu bar. If all module panels Inspector will display the data available at a se-
value.
are closed and the MeVisLab visual develop- lected module connector.
ment environment (IDE) has been minimized, The segmentation and quantification of the le-
the custom-designed UI of our image-process- sion involves the following steps:
ing application remains the only visible part. 1) Preprocessing: Thresholding and connected components
analysis to compute a mask of the lung image data.
Segmentation, Visualization, and 2) Segmentation: Region growing using an initial seed point.
Quantification of Lesions 3) Postprocessing: Dilation and computation of the border of the
The second example segments and computes the maximum lesion mask image.
diameter of a lung metastasis. Automatic segmentation meth- 4) Visualization: Direct volume rendering of the image data and
ods are often restricted to a specific type of object and data highlighting of the lesion.
on which they operate because of need for heuristics. Here, 5) Measurement: Quantification of the lesion’s maximum di-
the algorithm is based on the assumption that connected vas- ameter using the lesion mask image.
cular structures will be of small diameter when they finally Large parts of a chest CT image data will contain air voxels.
reach the lesion. Furthermore, the lesion must not infiltrate Air and tissue can be well distinguished in CT image data because
any connected tissue. The segmentation pipeline will be robust of their different voxel values. We use an IntervalThreshold
enough to ignore connections of the lesion to tissue and will module with a threshold value of −200 HU for the separation
also ignore connected vessel systems, in this case the pulmo- since air voxel values are around −1,000 HU and tissue values are
nary artery and vein. more than 0 HU. Separation of the lesion from the connected tis-
To obtain a first look onto the example in MeVisLab, please sue is more challenging; the voxel values may not differ enough
open the network file Lesion Segmentation.mlab [6]. In to prevent the region growing from spreading from the lesion
the left 2-D view, scroll to slice number 46 using the scroll into the adjacent regions. Since we know that the lesion will be
wheel of your mouse while pointing at the 2-D view. Place inside the lung, we can use a mask of the lung volume to cut off
a marker into the lesion as seen in Figure 7. The segmented the lesion at the border. This brings us the task of calculating a
lesion will be instantly highlighted in the 2-D as well as in lung volume mask.
the 3-D views. To display the maximum diameter of the le- We already obtained a mask of all nontissue voxels from
sion, open the module panel of the MaxDistance module. the IntervalThreshold module. Assuming the lung will
66 IEEE PULSE ▼ NOVEMBER/DECEMBER 2011
be the single largest object in the images, an assumption the size of a specific lesion, we opt for an interactive selection.
truly holding for a chest CT acquisition, a connected compo- Hence, the RegionGrowing module in the image pipeline
nent analysis using the ConnectedComponents module will use a marker, placed on a 2-D slice, to narrow down the im-
will find this single largest cluster. The resulting mask im- age mask to just the selected lesion.
age of this cluster will still contain holes and notches from The calculation of the diameter is a perfect example
the blood vessels, bronchial tubes, free-standing as well as of using the large image processing library of MeVisLab.
wall-attached lesions of the lung. We are particularly inter- There is no module that computes the maximum diameter
ested in the boundary of the volume, in the convex hull of of a 3-D image object from a given mask image. One could
the lung. The ConvexHull module computes the convex decide to implement such a module. However, MeVisLab
hull for voxels of a specific value. Now that we possess a mask provides a module to calculate the maximum distance be-
of tissue plus lesion and convex hull of the lung, we create tween a set of markers called XMarkerListMaxDis-
a new mask of all nonair voxels that are con- tance. Fortunately, we can convert a mask
tained within the lung using a Mask module. image into a set of markers using the module
The resulting mask will still contain vascular MaskToMarkers. However, applying this
structures. In contrast to the lesions, vascular Automatic module directly to our lesion mask image
structures are of elongated shape, a shape we segmentation would create many inside markers, while all
eliminate from the images using a fundamen- methods are often we need are markers on the outer border of
tal, morphological operation called opening. restricted to a specific the lesion. The Surround module offers a
Opening uses a structural element, called the mode to calculate just the boundary of a set
type of object and
kernel, to eliminate structures smaller than of voxels. Placing this module between the
this structural element. Hence, the kernel size
data on which they RegionGrowing and MaskToMarkers
parameter in the FastMorphology module operate because of modules saves computation time and en-
needs to be as large as the largest structures we need for heuristics. ables us to overlay the resultant border im-
would like to remove. The kernel will have a age onto the original image in the 2-D view
global effect on the mask image and also mod- to provide visual feedback about the seg-
ify the morphology of lesions; borders will be smoother and mentation to the user.
small notches will disappear. This is something to keep in Strictly speaking, the 3-D visualization of the lesion is only
mind when applying these operations. of limited diagnostic advantage. It provides an additional means
If there is only one lesion in the data set, we can stop now. for the inspection of the segmentation result, hence the cal-
The mask image will only contain this lesion. However, more of- culation of the maximum diameter. However, depending on
ten than not, the images will reveal multiple lesions. To quantify the chosen therapy, which could, for instance, perform local
3DView
SoCustomExaminerViewer
Measurement
MaxDistance SoBackground SoGVRTagVolume SoMLLUT SoGVRVolumeRenderer SoOrientationInset
XMarkerListMaxDistance
Dilation
RegionGrowing LUTConcat SoOrientationModel
MaskToMarkers FastMorphology
Opening
Postprocessing
Surround FastMorphology
ChangeLUTColor Visualization
2DView
View2D
Mask SoLUTEditor
SoView2DOverlay
ConvexHull
MarkerEditor IntervalThreshold
SoView2DMarkerEditor
ConnectedComponents
Segmentation Preprocessing
Invert
LocalImage Arithmetic1
FIGURE 7 The image-processing network used to quantify the maximum diameter of a wall-connected lesion in the lung.
NOVEMBER/DECEMBER 2011 ▼ IEEE PULSE 67
FIGURE 8 The final UI of the lesion quantification example.
destruction of the tumor, knowledge about the lesion’s sur- capabilities of human beings. For a human mind, an image
roundings could prove beneficial. represents a complex web of interrelated objects at different
Direct volume rendering is well suited for the 3-D depiction scales. It is prior knowledge on the characteristic properties
of this kind of data. Since air and tissue are represented by dif- and mutual relations of these image objects that enables a hu-
ferent values in CT and MR images, visual separation does not man to understand the image content. For a computer, an
need an explicit mask or any additional parameters for the image is merely a rectangular grid of voxels with intensity
transfer function (e.g., derived values, such as the gradient values. Therefore, most image-processing algorithms are lim-
or curvature). Highlighting the lesion itself is different. We ited to the evaluation of voxel values in local neighborhoods.
employ the slightly enlarged lesion mask image to apply a dif- To overcome this limitation, a rich set of tools for analyz-
ferent transfer function to the voxel values of the lesion in the ing images on the basis of objects has been created using the
original image. Voxel values with a mask value MeVisLab platform, similar to the way a hu-
of zero are rendered using the first transfer man understands an image. This set compris-
function, designed using the SoLUTEditor Visualization and es simple-to-use functionality for extracting
module, whereas the second transfer function spectral, shape, and relational object properties
is applied to voxel values with a mask value
analysis often involve from the image and for the knowledge-based
of one. This transfer function is derived from a separation evaluation of this information to identify the
the first by means of the ChangeLUTColor of structures. structures of interest.
module. It overrides just the color values of the In many areas of medical imaging, object-
transfer function with a highlighting color. The based image analysis solves problems that are
volume rendering itself is implemented in the SoGVRVolu- unsolvable by ordinary voxel-based image analysis. Complex
meRenderer module, which on its own offers various pa- image structures often do not become instantly obvious. In-
rameters to tune the visualization. The final UI of our lesion stead, image understanding is an iterative process of deriving
segmentation example can be seen in Figure 8. information from the image and applying prior knowledge.
This iterative process is demonstrated by an application for
Advanced Image Processing the detection of the spine in CT images [Figure 9(a)]. Voxel-
In this tutorial, we could only scratch the surface of medical im- based image-processing algorithms fail in this application
age processing. Two more advanced image-processing topics will because the spine does not exhibit unique intensity values.
conclude this tutorial and hopefully spark some interest in this Also, many are too distorted or inconspicuous to be iden-
field. There is still a lot to explore. tified in isolation. Object-based image analysis solves this
problem by exploiting prior knowledge that the spine is a
Object-Based Image Analysis loose chain of vertically aligned vertebrae with characteristic
Even the most sophisticated image analysis methods available spectral and shape properties. A watershed-based overseg-
today cannot cope with the remarkable pattern-recognition mentation creates the initial set of objects [Figure 9(b)]. The
68 IEEE PULSE ▼ NOVEMBER/DECEMBER 2011
(a) (b) (c) (d) (e)
FIGURE 9 Object-based detection of the spine: (a) original image, (b) initial oversegmentation, (c) detection of certain vertebrae, (d)
iterative classification of adjacent objects leading to full spine detection, and (e) refinement to close possible gaps.
V1 (%)
50
48 3.4
46 3.3
44 3.2
42 3.1
40 3
38 2.9
36 2.8
34 2.7
32 2.6
30 2.5
ROI_0 2.4
Ratio (t )
28
2.3
26
2.2
24 2.1
22 2
20 1.9
18 1.8
16 1.7
14 1.6
12 1.5
10 1.4
8 1.3
ROI_0
6 1.2
1.1 ROI_0(Fitted)
4
1
2 0 0.5 1 1.5 2 2.5 3 3.5 4 4.5
0 T (min)
(a) (b)
FIGURE 10 (a) Imaging the vasculature of a tumor with the help of a contrast agent. (b) The distribution of the contrast agent is
measured periodically over time (red crosses), and afterward a model curve (blue curve) is fitted to the measured data. The model
provides vascular information on the tissue and is superimposed onto the anatomical image data of the suspicious region as a
color map.
NOVEMBER/DECEMBER 2011 ▼ IEEE PULSE 69
first step of the analysis is to identify a few of these objects for the ROI. An example is shown in Figure 10, displaying
as vertebrae by employing strong constraints on their spec- the results for a very commonly used method—the dynamic
tral and shape properties [Figure 9(c)]. Adjacent vertebrae, contrast-enhanced magnetic resonance imaging (DCE-MRI)—
whose visual properties alone do not allow a certain classi- where a paramagnetic contrast agent is used to evaluate the
fication, can now be identified by their relative location to vasculature of tumors.
already detected vertebrae. Repeating the last step until the MeVisLab allows the user to import series of volumes
whole spine is recognized yields a robust detection method acquired over time of a number of medical imaging mo-
that even works with noisy images and vertebrae affected by dalities, for instance MRI, CT, and PET. There are modules
lesions [Figure 9(d)]. Remaining gaps (due to an inappropri- available to select an ROI in the images and generate time–
ate object presegmentation) can be closed with a subsequent intensity curves for these regions. Figure 10 shows an ex-
refinement step [Figure 9(e)]. ample evaluation created with MeVisLab. In Figure 10(a),
While we usually possess a wealth of knowledge about a curve generated from the DCE-MRI data is displayed. It
the objects in an image, it is often difficult to put this knowl- shows the temporal course of the ratio of signals between
edge into words. This applies, particularly, the contrast-agent-free first time period and
when the knowledge has to be stated numeri- time in which the contrast agent has arrived
cally, for instance, as the expected mean in- A series of DICOM in the tissue. The ratio is proportional to the
tensity or size of certain image objects. For images, which is the contrast agent concentration and can be used
this reason, our object-based image analysis most basic form for to optimize the parameters of the model de-
methods are capable to learn. All the user has scribing this time course. In this case, the
medical images, can
to do is to provide examples of relevant im- model describes the time course of the ratio
age objects, from which the computer then be retrieved from in dependence on some basic model parame-
derives the necessary knowledge to identify a disk using the ters: the vessel permeabilities and interstitial
further objects automatically. DirectDicom Import volume. The parameters are determined for
module. all voxels in a dynamic data set by optimiz-
Dynamic Imaging ing the model to the data using a least-square
Today, medical imaging offers a wide variety of algorithm. After conversion by a color map,
functional imaging that can give information on human tissue these parameters can be superimposed on top of the origi-
beyond that on shape and structure available from the imag- nal image data as shown in Figure 10.
ing information alone. These functional imaging processes can
yield valuable parameters on blood supply and circulation, and Acknowledgments
oxygen consumption or cell metabolism. The parameters find The authors wish to thank Dieter Haemmerich for fruitful
a broad application in diagnosis and treatment monitoring of discussions on the structure and contents of the tutorial and
various diseases. the MeVisLab team at MeVis Medical Solutions for exceptional
Frequently, these parameters are obtained by acquiring support.
subsequent images over time after injecting a contrast agent
into the bloodstream of the patient. Depending on the physi- Felix Ritter, André Homeyer, Hendrik Laue, Michael Schwier,
cal and biological properties of the contrast agents, the en- and Heinz-Otto Peitgen are with Fraunhofer MEVIS—Institute for
hancement course and pattern of the contrast agent will yield Medical Image Computing. Tobias Boskamp and Florian Link are
information on tissue properties. For example, Fludeoxyglu- with MeVis Medical Solutions AG.
cose-18 is processed in cells in the same manner as glucose
and therefore, is a good marker for cell metabolism. This can References
be exploited to measure the metabolism of tumors and dif- [1] MeVisLab. [Online]. Available: http://www.mevislab.de/
ferentiate tumor tissue from healthy tissue. Other contrast [2] R. C. Gonzalez and R. E. Woods, Digital Image Processing, 3rd ed.
agents are kept confined in the blood vessels and therefore, Englewood Cliffs, NJ: Prentice-Hall, 2007
their temporal and spatial distribution is related to blood vol-
[3] A. P. Dhawan, H. K. Huang, and D.-S. Kim, Principles and Ad-
ume and flow.
vanced Methods in Medical Imaging and Image Analysis, 1st ed.
Once data are acquired, the time-to-signal-curve can be
Singapore: World Scientific, 2008.
derived from the data, either by recording the signal changes
over time for single voxels or averaging over regions of interest [4] C. D. Hansen and C. R. Johnson, The Visualization Handbook.
(ROI) to obtain a mean time-to-signal curve. Each curve then New York: Elsevier, 2005.
can be evaluated using signal-processing techniques. Com- [5] A. Homeyer, M. Schwier, and H. K. Hahn, A generic concept for
monly used are compartment models describing the processing object-based image analysis,” in Proc. Int. Conf. Computer Vision
or distribution of the contrast agent by the tissue in the voxel Theory and Applications, 2010, pp. 530–533.
or ROI. The results of this analysis can either be a set of param- [6] F. Ritter. Example data of the article. [Online]. Available: http://
eter maps displaying the derived parameters for each voxel or, www.mevislab.de/IEEE_Pulse/
in the ROI-based version, a table containing the parameters
70 IEEE PULSE ▼ NOVEMBER/DECEMBER 2011
Das könnte Ihnen auch gefallen
- Unesco 97Dokument21 SeitenUnesco 97Leonell Romero BazanNoch keine Bewertungen
- Industrial X-Ray Computed TomographyVon EverandIndustrial X-Ray Computed TomographySimone CarmignatoNoch keine Bewertungen
- The CT Handbook Optimizing Protocols For Today's Feature Rich Scanners Chapter 1Dokument35 SeitenThe CT Handbook Optimizing Protocols For Today's Feature Rich Scanners Chapter 1Abdullah AliNoch keine Bewertungen
- Textbook of Urgent Care Management: Chapter 35, Urgent Care Imaging and InterpretationVon EverandTextbook of Urgent Care Management: Chapter 35, Urgent Care Imaging and InterpretationNoch keine Bewertungen
- Analysis of CT and MRI Image Fusion Using Wavelet TransformDokument5 SeitenAnalysis of CT and MRI Image Fusion Using Wavelet Transform18onaldo20orresNoch keine Bewertungen
- High Performance Deformable Image Registration Algorithms for Manycore ProcessorsVon EverandHigh Performance Deformable Image Registration Algorithms for Manycore ProcessorsNoch keine Bewertungen
- A Novel Approach To Multi Modal Hybrid Image Fusion Using Wavelet and Contourlet Transform For Medical Diagnosis ApplicationsDokument7 SeitenA Novel Approach To Multi Modal Hybrid Image Fusion Using Wavelet and Contourlet Transform For Medical Diagnosis ApplicationsRatnakarVarunNoch keine Bewertungen
- CT ScanDokument12 SeitenCT ScanNita ThrenitatiaNoch keine Bewertungen
- Image Fusion For Conformal Radiation TherapyDokument13 SeitenImage Fusion For Conformal Radiation TherapyAntonio López MedinaNoch keine Bewertungen
- DI Chap1Dokument9 SeitenDI Chap1HuntingparxxNoch keine Bewertungen
- Lecture No. 9 Basic Principles of CT ScanDokument24 SeitenLecture No. 9 Basic Principles of CT ScanMohammed Khalil SaeedNoch keine Bewertungen
- (IJCT-V2I1P13) Author: Mr. Roshan P. Helonde, Prof. M.R. JoshiDokument5 Seiten(IJCT-V2I1P13) Author: Mr. Roshan P. Helonde, Prof. M.R. JoshiIjctJournalsNoch keine Bewertungen
- Biomedical Image ProcessingDokument15 SeitenBiomedical Image ProcessingMokshada YadavNoch keine Bewertungen
- Lecture 2// 8.4 CT Numbers, Hounsfield Unit, Gray Scale and Image QualityDokument7 SeitenLecture 2// 8.4 CT Numbers, Hounsfield Unit, Gray Scale and Image QualityThome JerkNoch keine Bewertungen
- Computed TomographyDokument4 SeitenComputed TomographyemilyNoch keine Bewertungen
- Introduction To Medical ImagingDokument8 SeitenIntroduction To Medical ImagingTaj AlBarakatNoch keine Bewertungen
- Texture Analysis ReviewDokument28 SeitenTexture Analysis ReviewAnonymous 6WeX00mx0Noch keine Bewertungen
- AssignmentDokument3 SeitenAssignmentSuyash Maurya SuyashNoch keine Bewertungen
- Computed Tomography and Magnetic Resonance ImagingDokument14 SeitenComputed Tomography and Magnetic Resonance ImagingChecko LatteNoch keine Bewertungen
- Nugzar LipartelianiDokument78 SeitenNugzar LipartelianiTarun SinghNoch keine Bewertungen
- Module 1 Computed Tomography and Principles of OperationsDokument6 SeitenModule 1 Computed Tomography and Principles of OperationsWayne De Vergara PalaypayonNoch keine Bewertungen
- Wavelet Based Noise Reduction in CT-Images Using Correlation AnalysisDokument20 SeitenWavelet Based Noise Reduction in CT-Images Using Correlation Analysisprashant jadhavNoch keine Bewertungen
- Review Bio Medical ImagingDokument23 SeitenReview Bio Medical ImagingSweta TripathiNoch keine Bewertungen
- Document 39Dokument31 SeitenDocument 392022819384Noch keine Bewertungen
- Orden 2012 NeuroImageDokument9 SeitenOrden 2012 NeuroImageMichaelNoch keine Bewertungen
- Features: Medical Image RegistrationDokument4 SeitenFeatures: Medical Image RegistrationWill SmithNoch keine Bewertungen
- Simple Physical Principles and Medical Uses of Computed Tomography (CT) Scan A New AssessmentDokument10 SeitenSimple Physical Principles and Medical Uses of Computed Tomography (CT) Scan A New AssessmentIJAR JOURNALNoch keine Bewertungen
- T2 NanthagopalDokument10 SeitenT2 NanthagopalbelajartvkuNoch keine Bewertungen
- Cone Beam Technology: A Brief Technical: History of CAT and CBCTDokument9 SeitenCone Beam Technology: A Brief Technical: History of CAT and CBCTHugo MoralesNoch keine Bewertungen
- San 12193Dokument6 SeitenSan 12193Karla BFNoch keine Bewertungen
- Segmentation of Lung Nodule Using Image Processing Techniques From CT ImagesDokument4 SeitenSegmentation of Lung Nodule Using Image Processing Techniques From CT ImagesRachnaNoch keine Bewertungen
- The Generalized Contrast-to-Noise Ratio A Formal Definition For Lesion DetectabilityDokument15 SeitenThe Generalized Contrast-to-Noise Ratio A Formal Definition For Lesion DetectabilitydsfdsfNoch keine Bewertungen
- Slide 11 DiagimgDokument39 SeitenSlide 11 Diagimghayumbe08Noch keine Bewertungen
- Detection of Tumour in 3D Brain Image Using Mri Mining TechniquesDokument6 SeitenDetection of Tumour in 3D Brain Image Using Mri Mining TechniquesJeevan KishorNoch keine Bewertungen
- Welcome To International Journal of Engineering Research and Development (IJERD)Dokument6 SeitenWelcome To International Journal of Engineering Research and Development (IJERD)IJERDNoch keine Bewertungen
- EE4603 Ch1 (Intro)Dokument26 SeitenEE4603 Ch1 (Intro)ShuaiWangNoch keine Bewertungen
- Artefactos ConebeamDokument9 SeitenArtefactos ConebeamEmetac OdontológicoNoch keine Bewertungen
- Basic Physical Principles and Clinical ApplicationDokument17 SeitenBasic Physical Principles and Clinical ApplicationPopa Bogdan MihaiNoch keine Bewertungen
- Computed Tomography MachineDokument24 SeitenComputed Tomography MachineGaurav Molankar100% (1)
- Currently Available Maxillofacial CBCT EquipmentDokument4 SeitenCurrently Available Maxillofacial CBCT EquipmentzilniNoch keine Bewertungen
- Medical ImagingDokument9 SeitenMedical ImagingRajes WariNoch keine Bewertungen
- Tomography ThesisDokument7 SeitenTomography ThesisApril Smith100% (2)
- The Generation of DRR With Six ParametersDokument4 SeitenThe Generation of DRR With Six ParametersBojar DogNoch keine Bewertungen
- Part 2 Week 9 - Treatment PlanningDokument50 SeitenPart 2 Week 9 - Treatment PlanningdanNoch keine Bewertungen
- Mir DIP Report NarenDokument13 SeitenMir DIP Report Narenmatlab5903Noch keine Bewertungen
- Research Paper 1Dokument11 SeitenResearch Paper 1khizrullahkhanNoch keine Bewertungen
- Paper BRAIN TUMOR IMAGE CLASSIFICATION MODEL WITH EFFICIENTNETB3 (1909)Dokument5 SeitenPaper BRAIN TUMOR IMAGE CLASSIFICATION MODEL WITH EFFICIENTNETB3 (1909)Sathay AdventureNoch keine Bewertungen
- Module 1 Computed Tomography and Principles of OperationsDokument6 SeitenModule 1 Computed Tomography and Principles of OperationsWayne De Vergara PalaypayonNoch keine Bewertungen
- Predicting CT Image From MRI Data Through Feature Matching With Learned Nonlinear Local DescriptorsDokument11 SeitenPredicting CT Image From MRI Data Through Feature Matching With Learned Nonlinear Local DescriptorsANoch keine Bewertungen
- Volumetric 3D Display For Radiation Therapy PlanningDokument28 SeitenVolumetric 3D Display For Radiation Therapy PlanningNewberryNoch keine Bewertungen
- Image SegmentationDokument3 SeitenImage SegmentationrajasekarNoch keine Bewertungen
- Image Registration in Radiotherapy: Nilesh Kumar PG Radiation Physics Department of Radiation PhysicsDokument27 SeitenImage Registration in Radiotherapy: Nilesh Kumar PG Radiation Physics Department of Radiation Physicsnilesh kumarNoch keine Bewertungen
- Image Processing and Visualization TopicsDokument16 SeitenImage Processing and Visualization TopicsBradda Derru Nesta MarleyNoch keine Bewertungen
- An Efficient Visualization and Segmentation of Lung CT Scan Images For Early Diagnosis of CancerDokument5 SeitenAn Efficient Visualization and Segmentation of Lung CT Scan Images For Early Diagnosis of Cancermaruthi631Noch keine Bewertungen
- Virtual Simulation For Radiatiotherapy Treatment Using CT Medical DataDokument7 SeitenVirtual Simulation For Radiatiotherapy Treatment Using CT Medical Datasangeetha maniNoch keine Bewertungen
- T2 NanthagopalDokument10 SeitenT2 Nanthagopaladin80Noch keine Bewertungen
- CT Image Quality: Outsource, Inc. (All Rights Reserved)Dokument44 SeitenCT Image Quality: Outsource, Inc. (All Rights Reserved)bbkanilNoch keine Bewertungen
- Virtopsy Visualisation: Mixed Data Gradient Model For More Accurate Thin Bone Visualization in 3D RenderingDokument6 SeitenVirtopsy Visualisation: Mixed Data Gradient Model For More Accurate Thin Bone Visualization in 3D RenderingWolf SchweitzerNoch keine Bewertungen
- DTI HandbookDokument353 SeitenDTI HandbookIndicosa100% (1)
- A New Hybrid Approach For Brain Tumor Classification Using BWT-KSVMDokument6 SeitenA New Hybrid Approach For Brain Tumor Classification Using BWT-KSVMRasool ReddyNoch keine Bewertungen
- Curvelet-Based Multiscale Denoising Using Non-Local Means & Guided Image FilterDokument10 SeitenCurvelet-Based Multiscale Denoising Using Non-Local Means & Guided Image FilterRasool ReddyNoch keine Bewertungen
- Brain Tumor DatasetDokument17 SeitenBrain Tumor DatasetRasool ReddyNoch keine Bewertungen
- Satellite Image ClassificationDokument52 SeitenSatellite Image ClassificationRasool ReddyNoch keine Bewertungen
- GUI Implementation of Image Encryption and Decryption Using Open CVDokument4 SeitenGUI Implementation of Image Encryption and Decryption Using Open CVRasool ReddyNoch keine Bewertungen
- Adaptive Systems: Performance You Can SeeDokument8 SeitenAdaptive Systems: Performance You Can SeeRasool ReddyNoch keine Bewertungen
- Techniques and Antenna Radar Sys Anti Jam An D Methods For Adaptive Stem Polarization Optim ND Anti Clutter Applicati Single Mization For IonsDokument4 SeitenTechniques and Antenna Radar Sys Anti Jam An D Methods For Adaptive Stem Polarization Optim ND Anti Clutter Applicati Single Mization For IonsRasool ReddyNoch keine Bewertungen
- P S CK: Seshadri Rao Knowledge Village::GudlavalleruDokument2 SeitenP S CK: Seshadri Rao Knowledge Village::GudlavalleruRasool ReddyNoch keine Bewertungen
- Prasar Bharati 2013Dokument8 SeitenPrasar Bharati 2013Rasool ReddyNoch keine Bewertungen
- Journallistofscopus PDFDokument630 SeitenJournallistofscopus PDFSatyanarayana RentalaNoch keine Bewertungen
- DAY Wise ProgramsDokument61 SeitenDAY Wise ProgramsRasool ReddyNoch keine Bewertungen
- Resistors: Fig: 1.1 Colour Coding On ResistorDokument107 SeitenResistors: Fig: 1.1 Colour Coding On ResistorRasool ReddyNoch keine Bewertungen
- Cyclic Encoding & DecodingDokument3 SeitenCyclic Encoding & DecodingRasool ReddyNoch keine Bewertungen
- S.No - Project Title Name of The Students Area of Specialization PEO PODokument4 SeitenS.No - Project Title Name of The Students Area of Specialization PEO PORasool ReddyNoch keine Bewertungen
- Accessing Hardware: Code Club PihackDokument2 SeitenAccessing Hardware: Code Club PihackRasool ReddyNoch keine Bewertungen
- Real Time Vehicle Monitoring and Tracking System For School Bus Via Beagle BoneDokument4 SeitenReal Time Vehicle Monitoring and Tracking System For School Bus Via Beagle BoneRasool ReddyNoch keine Bewertungen
- Print Order Details4Dokument1 SeitePrint Order Details4Rasool ReddyNoch keine Bewertungen
- DSP Important Questions Unit-WiseDokument6 SeitenDSP Important Questions Unit-WiseRasool Reddy100% (4)
- Gudlavalleru Engineering College: Criteria 3.5 Process of Identifying Curricular Gaps To The Attainment of The Cos/PosDokument2 SeitenGudlavalleru Engineering College: Criteria 3.5 Process of Identifying Curricular Gaps To The Attainment of The Cos/PosRasool ReddyNoch keine Bewertungen
- Cmos Digital Ic Design Mid PaperDokument2 SeitenCmos Digital Ic Design Mid PaperRasool ReddyNoch keine Bewertungen
- On Performance Improvement of Wireless Push Systems Via Smart AntennasDokument5 SeitenOn Performance Improvement of Wireless Push Systems Via Smart AntennasRasool ReddyNoch keine Bewertungen
- Graphical Representation Uniform and Non Uniform MotionDokument24 SeitenGraphical Representation Uniform and Non Uniform MotionKashyap PatelNoch keine Bewertungen
- Applications of Lasers PDFDokument4 SeitenApplications of Lasers PDFRUPAM GHOSHNoch keine Bewertungen
- Tos Presentation DemoDokument6 SeitenTos Presentation DemopetyorkfiveNoch keine Bewertungen
- C5X Design Rules 45000099 RevTDokument89 SeitenC5X Design Rules 45000099 RevTMohammed TamheedNoch keine Bewertungen
- Corona Discharge - Wikipedia, The Free EncyclopediaDokument8 SeitenCorona Discharge - Wikipedia, The Free EncyclopediavagoliyoNoch keine Bewertungen
- 2309.04860 Approximation Results For Gradient Descent TrainedDokument69 Seiten2309.04860 Approximation Results For Gradient Descent TrainedhongnhNoch keine Bewertungen
- Lead Acid BatteryDokument11 SeitenLead Acid BatterymapalptsNoch keine Bewertungen
- Pioneer VenusDokument256 SeitenPioneer VenusBob Andrepont100% (2)
- Gas Insulated SwitchgearDokument33 SeitenGas Insulated SwitchgearAkinwande Quadri100% (2)
- 1.4 Measurements: Chapter 1 - Essential Ideas 29Dokument7 Seiten1.4 Measurements: Chapter 1 - Essential Ideas 29Urvi MohanNoch keine Bewertungen
- The Design of Seismic Reaction Block of BridgeDokument8 SeitenThe Design of Seismic Reaction Block of BridgeRoshan KejariwalNoch keine Bewertungen
- Acoustics - Determination of Occupational Noise Exposure - Engineering Method (ISO 9612:2009)Dokument15 SeitenAcoustics - Determination of Occupational Noise Exposure - Engineering Method (ISO 9612:2009)kalyan1990Noch keine Bewertungen
- Chapter 14 PolynomialsDokument20 SeitenChapter 14 PolynomialsOzzy PingBoi100% (2)
- 9701 Nos Ps 2Dokument6 Seiten9701 Nos Ps 2Hubbak Khan0% (1)
- LAS No6Dokument6 SeitenLAS No6Justine FaustoNoch keine Bewertungen
- 1502 04177Dokument380 Seiten1502 04177Anju SharmaNoch keine Bewertungen
- Urea ProjectDokument31 SeitenUrea Projectaman singhNoch keine Bewertungen
- 2080iq4 2Dokument2 Seiten2080iq4 2RajeshNoch keine Bewertungen
- Quasi-3D Modelling of Two-Phase Slug Flow in PipesDokument12 SeitenQuasi-3D Modelling of Two-Phase Slug Flow in PipeskumarNoch keine Bewertungen
- Chapter 06 PRBX00Dokument8 SeitenChapter 06 PRBX00ausgrumb xoneNoch keine Bewertungen
- C FIX Report0Dokument7 SeitenC FIX Report0Laurence SarmientoNoch keine Bewertungen
- Surge Propagation in Multiconductor Transmission Lines Below GroundDokument42 SeitenSurge Propagation in Multiconductor Transmission Lines Below GroundDejanNoch keine Bewertungen
- IEEE-A Primer On Capacitor Bank Protection PDFDokument6 SeitenIEEE-A Primer On Capacitor Bank Protection PDFGustavo AguayoNoch keine Bewertungen
- Deformable Bodies (Thermal Stress)Dokument11 SeitenDeformable Bodies (Thermal Stress)Kristelle GinezNoch keine Bewertungen
- Resume 2Dokument4 SeitenResume 2Kutty166Noch keine Bewertungen
- Bulk Deformation ProcessesDokument71 SeitenBulk Deformation ProcessesHavenesh HaveNoch keine Bewertungen
- Easy ReflectDokument16 SeitenEasy Reflectali kundiNoch keine Bewertungen
- Quotation Yoke WIFL Sanaswadi 201718Dokument4 SeitenQuotation Yoke WIFL Sanaswadi 201718Deipak HoleNoch keine Bewertungen
- 1750 22Dokument4 Seiten1750 22hogoyoNoch keine Bewertungen
- The Innovators: How a Group of Hackers, Geniuses, and Geeks Created the Digital RevolutionVon EverandThe Innovators: How a Group of Hackers, Geniuses, and Geeks Created the Digital RevolutionBewertung: 4 von 5 Sternen4/5 (331)
- The Journeyman Electrician Exam Study Guide: Proven Methods for Successfully Passing the Journeyman Electrician Exam with ConfidenceVon EverandThe Journeyman Electrician Exam Study Guide: Proven Methods for Successfully Passing the Journeyman Electrician Exam with ConfidenceNoch keine Bewertungen
- The Innovators: How a Group of Hackers, Geniuses, and Geeks Created the Digital RevolutionVon EverandThe Innovators: How a Group of Hackers, Geniuses, and Geeks Created the Digital RevolutionBewertung: 4.5 von 5 Sternen4.5/5 (543)
- Foundations of Western Civilization II: A History of the Modern Western World (Transcript)Von EverandFoundations of Western Civilization II: A History of the Modern Western World (Transcript)Bewertung: 4.5 von 5 Sternen4.5/5 (12)
- The Homeowner's DIY Guide to Electrical WiringVon EverandThe Homeowner's DIY Guide to Electrical WiringBewertung: 5 von 5 Sternen5/5 (2)
- Practical Troubleshooting of Electrical Equipment and Control CircuitsVon EverandPractical Troubleshooting of Electrical Equipment and Control CircuitsBewertung: 4 von 5 Sternen4/5 (5)
- Electrical Engineering 101: Everything You Should Have Learned in School...but Probably Didn'tVon EverandElectrical Engineering 101: Everything You Should Have Learned in School...but Probably Didn'tBewertung: 4.5 von 5 Sternen4.5/5 (27)
- Beginner's Guide to Reading Schematics, Fourth EditionVon EverandBeginner's Guide to Reading Schematics, Fourth EditionBewertung: 3.5 von 5 Sternen3.5/5 (10)
- Arduino and Raspberry Pi Sensor Projects for the Evil GeniusVon EverandArduino and Raspberry Pi Sensor Projects for the Evil GeniusNoch keine Bewertungen
- Cleanroom Technology: Fundamentals of Design, Testing and OperationVon EverandCleanroom Technology: Fundamentals of Design, Testing and OperationNoch keine Bewertungen
- 2022 Adobe® Premiere Pro Guide For Filmmakers and YouTubersVon Everand2022 Adobe® Premiere Pro Guide For Filmmakers and YouTubersBewertung: 5 von 5 Sternen5/5 (1)
- Practical Guide to FMEA : A Proactive Approach to Failure AnalysisVon EverandPractical Guide to FMEA : A Proactive Approach to Failure AnalysisBewertung: 5 von 5 Sternen5/5 (1)
- Marine Rudders and Control Surfaces: Principles, Data, Design and ApplicationsVon EverandMarine Rudders and Control Surfaces: Principles, Data, Design and ApplicationsBewertung: 4.5 von 5 Sternen4.5/5 (3)
- Electronics All-in-One For Dummies, 3rd EditionVon EverandElectronics All-in-One For Dummies, 3rd EditionBewertung: 5 von 5 Sternen5/5 (2)
- Programming the Raspberry Pi, Third Edition: Getting Started with PythonVon EverandProgramming the Raspberry Pi, Third Edition: Getting Started with PythonBewertung: 5 von 5 Sternen5/5 (2)
- Empires of Light: Edison, Tesla, Westinghouse, and the Race to Electrify the WorldVon EverandEmpires of Light: Edison, Tesla, Westinghouse, and the Race to Electrify the WorldBewertung: 4 von 5 Sternen4/5 (87)
- Guide to the IET Wiring Regulations: IET Wiring Regulations (BS 7671:2008 incorporating Amendment No 1:2011)Von EverandGuide to the IET Wiring Regulations: IET Wiring Regulations (BS 7671:2008 incorporating Amendment No 1:2011)Bewertung: 4 von 5 Sternen4/5 (2)
- Open Radio Access Network (O-RAN) Systems Architecture and DesignVon EverandOpen Radio Access Network (O-RAN) Systems Architecture and DesignNoch keine Bewertungen
- Practical Electrical Wiring: Residential, Farm, Commercial, and IndustrialVon EverandPractical Electrical Wiring: Residential, Farm, Commercial, and IndustrialBewertung: 3.5 von 5 Sternen3.5/5 (3)
- The Fast Track to Your Extra Class Ham Radio License: Covers All FCC Amateur Extra Class Exam Questions July 1, 2020 Through June 30, 2024Von EverandThe Fast Track to Your Extra Class Ham Radio License: Covers All FCC Amateur Extra Class Exam Questions July 1, 2020 Through June 30, 2024Noch keine Bewertungen