Beruflich Dokumente
Kultur Dokumente
Ambion IsoRNA Protocol S09update
Hochgeladen von
Mohammad El BastOriginalbeschreibung:
Copyright
Verfügbare Formate
Dieses Dokument teilen
Dokument teilen oder einbetten
Stufen Sie dieses Dokument als nützlich ein?
Sind diese Inhalte unangemessen?
Dieses Dokument meldenCopyright:
Verfügbare Formate
Ambion IsoRNA Protocol S09update
Hochgeladen von
Mohammad El BastCopyright:
Verfügbare Formate
Cell and Molecular Biology Laboratory BIOL 406
Isolation of Total RNA from Yeast
(This protocol revised&adapted from Dr. Karen Bernd, at Davidson College, N.C.)
Objectives: 1. To learn how to work with yeast and extract Total RNA from them. 2. Understand the importance of RNases and how to protect RNA samples. 3. Quantitate and check purity of nucleic acid samples 4. Understand the difference between fermentation and respiration in yeast 5. Understand the steps required in experimental design and planning
READ THIS PROTOCOL CAREFULLY. MAKE SURE YOU ARE PREPARED TO ASK ANY TECHNICAL OR THEORETICAL QUESTIONS BEFORE WE BEGIN THIS EXPERIMENT. IF IN DOUBT ABOUT ANYTHING, ASK!
Isolation of total RNA from yeast
RNA expression is one way of measuring gene activity. As genes are turned on, mRNA levels for those genes increase, generally resulting in increased protein expression. While cells have many ways of regulating proteins, differences in the functional state of the cell correlates directly with changes in mRNA levels. The cells use these new proteins to respond to the current environmental challenges within the cell. In this experiment, we will monitor the yeasts response to decreasing concentrations of glucose in the media as the yeast are forced to live under less favorable conditions. In rich media (i.e. high in glucose) the yeast metabolize the sugars to ethanol through anaerobic fermentation, but under less favorable conditions (little or no glucose) , they start metabolizing ethanol by aerobic respiration. This process of changing food sources in yeast is called the diauxic shift. During the diauxic shift the yeast undergo substantial changes in gene expression related to the many biochemical pathways involved in the different methods of metabolism. We will try to gain insights into gene expression by examining total mRNA levels in yeast at critical time points throughout the shift using microarray technology. The first step
Cell and Molecular Biology Laboratory BIOL 406
in this process is to obtain yeast cultures at time points that represent the changes that occur during the diauxic shift, starting with yeast that have been grown in the presence of high glucose (i.e. the positive control) and subsequently in lower glucose concentrations. Your job is to isolate the total RNA at one time point. We will share our data, to get a complete time course of the diauxic shift when we complete the microarray analysis for each time point later in the semester. In order to characterize the diauxic shift in yeast, you will be isolating total RNA from cultures of YEAST (STRAIN DBY10009) a
generous gift from the Botstein Lab, Princeton University, grown for different lengths of time. This particular strain of yeast has been completely
sequenced. Later on, we will analyze gene expression of the ~6000 genes in yeast and try to correlate differences with their growing conditions. Your results for the future microarray work will depend on how well you complete this initial phase of the experiment. Make sure you work CAREFULLY. The most important step in RNA isolation is to remove as many sources of RNases from your work area as possible. RNases are enzymes whose entire purpose is to degrade RNA. They are very stable, low molecular weight proteins that can withstand high temperature and they are EVERYWHERE (yes, be paranoid!) ANY contamination with RNase will destroy your sample and pretty much wreck your day, experimentally speaking. RNA is extremely sensitive to degradation by RNases. How carefully you handle your samples and transfer solutions will have a huge impact on your yield. You must get rid of RNases. Your objective is to isolate a large quantity (perhaps 100 whole micrograms) of high molecular weight, undegraded total RNA. Read all instructions before touching anything. Make sure you have cleaned your bench space before beginning. Extra caution now will save you much work later. The main source of RNase contamination is your hands. Since it is out of the question to bake your hands at 210 for 15 hours to
Cell and Molecular Biology Laboratory BIOL 406
inactivate the enzyme, you must do everything you can to eliminate contact between your skin, things your skin has touched, and your precious samples. Clear your bench of all but the bare essentials. Once you have broken open the cells the RNAs are released and are susceptible to degradation by RNAases. Wear gloves and wash down your entire area with 'RNase ZAP' (special detergent that helps control RNases a bit.) All microfuge tubes and pipet tips have only been touched with gloves and have been sterilized extensively. The protocol is not difficult but you must be organized and careful.
Cell and Molecular Biology Laboratory BIOL 406
A. Materials:
Yeast (Saccharomyces cerevisiae Strain DBY10009) YPD Media (autoclaved) 10 g/L Yeast Extract 20 g/L Peptone 20 g/L Dextrose Lyticase (20 Units/mg) (store @ 4C, incubate @ 35C) (lyticase is an enzyme that breaks down yeast cells walls) BME (-mercaptoethanol) RNAse ZAP 100% ethanol
Use ACS grade 100% ethanol or better. You will need 40 mL to add to the Wash Solution 2/3 Concentrate, and 1.25 mL per sample for use in step D.3.
15 ml tubes with tight fitting lids (ie corning) 50 ml Conical tubes 1.6 ml Eppendorf tubes 200 ul & 1000 ul RNase Free pipet tips Pipetmen
1.2 M Sorbitol, 10 mM KPhos pH 7.2
Potassium Phosphate (mw = 136.09 g/mol) Sorbitol buffer (mw = 182.17 g/mol)
Ambion RiboPure Yeast Kit (Cat #AM1926)
Ambion RiboPure-Yeast is a rapid RNA isolation kit that combines disruption of yeast cells with Zirconia Beads, phenol extraction of the lysate, and glass-fiber filter purification of the RNA. It can be used to isolate total RNA from a variety of yeast species. RiboPure-Yeast was extensively tested with Saccharomyces cerevisiae, Schizosaccharomyces pombe, Pichia pastoris,and Candida albicans. Amount
50 50 100 24 mL 2.4 mL 24 mL 95 mL 35 mL 50 mL
Component
1.5 mL Screw Cap Tubes Filter Cartridges Collection Tubes (2 mL) Lysis Buffer 10% SDS* Phenol:Chloroform:IAA Binding Buffer Wash Solution 1* Wash Solution 2/3 Concentrate
Storage
room temp room temp room temp 4C 4C 4C 4C 4C 4C
Cell and Molecular Biology Laboratory BIOL 406
Add 40 mL 100% ethanol before use 5 mL 80 g 550 L 200 L 550 L Elution Solution Zirconia Beads (0.5 mm dia) 10X DNase 1 Buffer DNase 1 DNase Inactivation Reagent 4C 20C 20C 20C 20C
* If a precipitate develops during storage, dissolve the precipitate before use by warming the solution to 37C. Phenol:Chloroform:IAA is a poison and an irritant. Contact with it will cause burns. Use gloves and other personal protection equipment when working with this reagent. This reagent contains guanidinium thiocyanate, a potentially hazardous substance; use with appropriate caution.
Vortex Mixer w/ adapter (Cat Ambion #AM10024) 95C Heating Blocks filled with deionized water 30C & 37C Water Bath Incubators Swinging bucket centrifuge @ 4C (G-309) 1.2% Agarose Gels DNA Gel boxes 10X Nucleic Acid Sample loading dye HyperLadder I Wide Range Markers Gel Running Bufffer (1 liter electrophoresis buffer 1x TBE) Ethidium Bromide Power Supply
NOTES:
Lab bench and pipettors Before working with RNA, it is always a good idea to clean the lab bench and pipettors with an RNase decontamination solution (e.g. Ambion RNase Zap Solution). Gloves and RNase-free technique Wear laboratory powder free gloves at all times during this procedure and change them frequently. They will protect you from the reagents, and they will protect the RNA from nucleases that are present on skin. Use RNase-free tips to handle the wash solutions and the Elution Solution, and NEVER put used tips into the kit reagents.
Cell and Molecular Biology Laboratory BIOL 406
Keep reagents on ice Always close pipet tip boxes Dispose of tips in waste containers, not on the bench Never use tips more than once! B. Preparing yeast cells by making spheroplasts
Yeast cells are grown in YPD media at 30 C in a shaking incubator. Yeast cultures grow exponentially until the media is exhausted. Growth of the yeast cultures is monitored by optical density, (Absorbance at 600 nm). Yeast are grown very similarly to bacteria. Like bacteria, they have different stages of growth. Scientists divide these stages of growth into early-log phase, mid log-phase, late log-phase, and the stationary phase. These phases are based on the amount of yeast that are in the broth and how quickly they are able to grow and divide based on nutrient use. Reading the absorbance of the sample at 600 nm is a way in which you can determine which phase of the yeast growth cycle you are in and how many cells you have. Here is a chart that will show you how this relates to yeast growth. You should record the O.D. of the class cultures.
OD 600 Early log-phase mid log-phase late log-phase stationary phase < 0.4 0.4 - 1.7 1.7 -6.6 > 6.6
cells/ml < 107 1-5 x 107 5 x107 - 2 x 108 8 2 x 108
Haploid yeast cells contain approximately 1.2 pg of RNA per cell.
Cell and Molecular Biology Laboratory BIOL 406
The volume of culture for each time point was adjusted so that each sample (time point) contains the same amount of yeast. The cultures were inoculated and grown for 9 hours, then samples were withdrawn from the culture every 4 hours. The yeast cells were pelleted and then flash frozen and stored at -80C. Your group will receive yeast cells that were grown for a certain period of time (9 hrs, 13 hrs, 17 hrs or 21 hrs). We expect gene expression in the yeast to change because glucose was depleted over the time course. Yeast contain a hard cell wall. To improve our RNA yields we need to breakdown the cell wall so that the cells may be lysed more easily. Spheroplasts are yeast cells where the cell wall has been enzymatically degraded (ie using Lyticase). 1) Each group receives a yeast cell pellet that has been collected at one of the above time points (9 hrs, 13 hrs, 17 hrs or 21 hrs). 2) Label your yeast sample according to the time point youre given so that you can identify your tube. Samples were standardized to contain 21 OD600 units. (i.e. 30 ml x 0.7 OD600 Absorbance units) Predict approximately how much RNA you should isolate. 3) Get your supplies ready: yellow and blue tips for pipetmen, 1.6 ml Microfuge tubes, and Potassium Phosphate/Sorbitol buffer. The next 3 steps are called a wash--they serve to move the frozen cells into a solution that is correctly buffered for the next procedure. 4) Add 1.0 ml of Potassium Phosphate/Sorbitol buffer to your yeast pellet. Let the yeast thaw on ice for at least 5 minutes. Resuspend the pellet by gently pipetting. 5) Label one 1.6 ml (large) eppendorf tube. Transfer the cells from the 15 ml conical tubes to the eppendorf tube. 6) Place the eppendorf tubes in the microcentrifuge (balancing tubes). Spin at 11,200g for 3min @ 4C.
Cell and Molecular Biology Laboratory BIOL 406
7) Pour off supernatant (without disturbing pellet). Remove any residual supernatant with a pipetman. Resuspend pellet in 1.0 ml Potassium Phosphate/Sorbitol buffer. 8) Take tubes to hood. Add: 3l -mercaptoethanol (a reducing agent that smells very bad) 320 l of lyticase (enzyme that degrades cell wall components) 9) Place tubes in the incubator at 30C (check TEMP!!) for 15min. The cells are now spheroplasts. Without a cell wall they are alive but structurally much weaker. Care must be taken so that the cells don't burst before you want them to. (How could the buffer they are in help keep them intact?) NOTE: yeast will repair their cell wall if the enzyme is removed so we cannot just keep a stock of 'wall free' yeast. You must now get the cells ready for the next set of steps by washing away the lyticase and -mercaptoethanol. 10) Spin in microcentrifuge at 5600 g. Remove supernatant by pipeting (discard in hood to contain the BME smell). 11) Add 500l potassium phosphate/sorbitol buffer. Repeat step 10 (one time). 12) While centrifuging make sure your ice bucket contains: 1 tube 1 tube 1 tube precipitate) 1 tube (screw cap) containing glass (zirconia) beads of Lysis Buffer of 10% SDS (Warm @ 37C to dissolve any of Phenol:Chloroform:IAA
Clean RNA isolation area with 'RNAse ZAP' solution
Cell and Molecular Biology Laboratory BIOL 406
(1 spray bottle for lab)
Isolating RNA
The RiboPure-Yeast method disrupts yeast cell walls by beating cells mixed with an aqueous lysis buffer, SDS, phenol and 0.5 mm Zirconia Beads on a vortex adaptor (e.g. Ambion Cat #AM10024) for 10 min. Up to 3 x 108 cells can be processed at a time. The lysate is centrifuged to separate the aqueous phase, which contains the RNA, from the lower organic phase, which contains proteins, polysaccharides, and other cellular debris. The aqueous lysate is diluted with Binding Buffer and ethanol, and is drawn through a glass-fiber filter, which immobilizes the RNA. Contaminants are washed from the filter, and finally the RNA is eluted in a low ionic strength solution. Residual DNA is removed by treating the RNA with Ambion DNAfree reagents (DNAse enzymes), included in the kit. As with all glass-fiber filter purification methods, 5S ribosomal RNAs and tRNAs are not quantitatively recovered using the RiboPure-Yeast Kit. The entire RNA isolation procedure and DNase treatment require approximately 1.5 hrs, starting with fresh, snap-frozen cultured yeast cells. The resulting RNA should be of superb quality, free of DNA and proteins, and suitable for use in virtually any downstream application.
FROM THIS POINT ON DO NOT TOUCH ANYTHING ON YOUR BENCH WITHOUT WEARING GLOVES. DO NOT 'DOUBLETOUCH' TIPS DO NOT WALK AROUND WITH OPEN TUBES DO NOT LAUGH, TALK, OR BREATHE OVER OPEN TUBES (To quote the movies "be afraid--be very afraid")
C. Cell Disruption and Initial RNA Purification
1. Resuspend cells in lysis reagents Add the following to your yeast cell pellet in the order shown, and resuspend by vortexing vigorously for 1015 seconds: (CAUTION: these reagent are extremely caustic; handle with respect) Amount Component 480 L Lysis Buffer 48 L 10% SDS (Check to make sure no precipitate is present) 480 L Phenol:Chloroform:IAA 6 2. Add mixture to Zirconia Beads Transfer the mixture of cells and lysis reagents to the prepared tube containing ~750 L cold Zirconia Beads, and securely fasten the lid.
Cell and Molecular Biology Laboratory BIOL 406
10
3. Beat cells for 10 min on vortex mixer with a vortex adapter Position the sample tubes horizontally on the vortex adapter with the tube caps towards the center. Turn the vortex mixer on at maximum speed and beat for 10 min to lyse the yeast cells. 4. Centrifuge the lysed cells for 5 min at room temp, then transfer the aqueous phase to the 15 mL tube that contains 1.9 ml of binding buffer as explained below. a. Centrifuge for 5 min at 16,100xg at room temp to separate the aqueous phase, containing the RNA, from the organic phase. b. Transfer the aqueous phase (top), containing the partially purified RNA, to the 15 mL tube labeled Binding Buffer. Typically, the recovered volume from the cell lysis will be approximately 530 L.
D. Final RNA Purification
1. Before you start: Check Wash Solution 1 and redissolve by heating to 37C if necessary A precipitate may form in the Wash Solution 1. Examine the reagents carefully before use, and if a precipitate is visible, redissolve it by heating the solution(s) to 37C, and agitating the tube occasionally. Preheat Elution Solution to ~95C Place 50 L Elution Solution per sample into the heat block set to 95100C. The preheated Elution Solution will be used in step D.8. Inspect the Filter Cartridges Briefly inspect your Filter Cartridge before use. Occasionally, the glass fiber filters may become dislodged during shipping. If this is the case, gently push the filter down to the bottom of the cartridge using the wide end of a RNase-free pipette tip. NOTE: All centrifugation steps should be done at an RCF of ~13,00016,000xg. This is typically near the maximum speed setting on a microfuge. 2. Your sample now has a volume of ~2.4 ml (~530L recovered from the lysis step and 1.9 ml of Binding Buffer.) This corresponds to 350 L Binding Buffer for each 100 L of partially purified RNA. 3. Add 1.25 mL 100% ethanol to your sample Add 1.25 mL 100% ethanol and mix thoroughly. (This corresponds to 235 L ethanol for each 100 L of partially purified RNA.)
Cell and Molecular Biology Laboratory BIOL 406
11
4. Draw sample through a Filter Cartridge a. Apply 700 L of the sample mixture to a Filter Cartridge assembled in a Collection Tube. The total volume of the lysate/ethanol mixture is ~3.7 mL, since the Filter Cartridge can only accommodate 700 L, you will need to draw the lysate mixture through the Filter Cartridge in several applications of 700 L at a time. b. Centrifuge for ~0.51 min or until the lysate/ethanol mixture is through the filter. c. Discard the flow-through and return the Filter Cartridge to the Collection Tube. d. Repeat as necessary to pass the entire sample through the Filter Cartridge. NOTE: The RNA is now bound to the filter in the Filter Cartridge. 5. Wash filter with 700 L Wash Solution 1 a. Wash the filter by adding 700 L Wash Solution 1 to the Filter Cartridge, and centrifuging for ~1 min or until all of the liquid is through the filter. b. Discard the flow-through and return the Filter Cartridge to the same Collection Tube. 6. Wash filter with 2x500 L Wash Solution 2/3 a. Wash the filter by adding 500 L Wash Solution 2/3 to the Filter Cartridge. Draw it through the filter as in the previous step. b. Repeat with a second 500 L aliquot of Wash Solution 2/3. 7. Centrifuge for 1 min to remove excess wash solution from the filter a. Centrifuge the Filter Cartridge for 1 min to remove excess wash. b. Transfer the Filter Cartridge to a fresh 2 mL Collection Tube. 8. Elute RNA in 2x25 L preheated Elution Solution a. Elute RNA by applying 25 L Elution Solution, preheated to 95100C, to the center of the filter. b. Centrifuge for 1 min. c. Repeat the elution step with a second 25 L aliquot of preheated Elution Solution into the same Collection Tube. Make sure your RNA is clearly labeled with your name, date, sample info etc. This a good stopping point, so we will freeze our samples until next week to finished the purification and analysis.
STOP and Store samples at -70 C --
Cell and Molecular Biology Laboratory BIOL 406
12
Continue here at the next lab.
E. DNase I Treatment
RNA isolated using RiboPure-Yeast will contain contaminating chromosomal DNA. To remove it, we recommend including this DNase 1 treatment. The DNase Treatment and Removal Reagents provided with the RiboPure-Yeast Kit include ultrapure DNase I and reaction buffer that are rigorously tested to be sure that no RNase is present. This procedure makes use of a simple process to inactivate the DNase without the risk incurred by heating the RNA, or the hassles of adding EDTA or extracting with phenol/chloroform. 1. Assemble the DNase digestion reaction Assemble the DNase digestion reaction at room temp. Amount 50 L 5 (1/10th vol) 4 L Component RNA sample (from step D.8) 10X DNase 1 Buffer (i.e. 510 L) DNase 1 (8 UNITS)
2. Incubate 30 min at 37C Incubate the DNase digestion reaction for 30 min at 37C. 3. Treat with 0.1 volume DNase Inactivation Reagent a. Resuspend the DNase Inactivation Reagent by flicking or vortexing the tube, then add 0.1 volume DNase Inactivation Reagent to each sample. For a typical 59 L DNase digestion reaction, use 6 L DNase Inactivation Reagent. b. Mix by vortexing and allow reactions to stand 5 min at room temp. c. Centrifuge 23 min at top speed in a microfuge (10,000 x g) at room temp to pellet the DNase Inactivation Reagent. Transfer the RNA (supernatant) to a fresh tube.
THIS IS YOUR RNA. Place the tube on ice!
Cell and Molecular Biology Laboratory BIOL 406
13
Your RNA should always be stored on ice or frozen to slow degradation (why would cold slow degradation?) Question: What types of RNA have you just isolated? (how many different kinds of RNA are in a cell?)
F. RNA Storage
RNA samples can be stored at 20C for up to 12 months, or at 80C for longer than 2 months.
Checking RNA Integrity
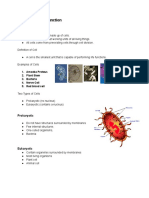
The quality of your RNA can be checked by agarose gel electrophoresis Prepare a 5 ug sample of your RNA for analysis by 1.2% Agarose gel. Coordinate with other groups to load samples (one sample from each time point) and markers on the gel. Run the gel at 120 mV for approximately 20 minutes until the two markers divide the gel into roughly equal thirds Stain the gel with ethidium bromide. Note the position of bands on the gel and the intensity of the staining. (Does your sample look like the ones shown the sample gel?)
1 = Degraded RNA 2 = Good RNA 3 = Good RNA
Das könnte Ihnen auch gefallen
- Shoe Dog: A Memoir by the Creator of NikeVon EverandShoe Dog: A Memoir by the Creator of NikeBewertung: 4.5 von 5 Sternen4.5/5 (537)
- Never Split the Difference: Negotiating As If Your Life Depended On ItVon EverandNever Split the Difference: Negotiating As If Your Life Depended On ItBewertung: 4.5 von 5 Sternen4.5/5 (838)
- Elon Musk: Tesla, SpaceX, and the Quest for a Fantastic FutureVon EverandElon Musk: Tesla, SpaceX, and the Quest for a Fantastic FutureBewertung: 4.5 von 5 Sternen4.5/5 (474)
- The Subtle Art of Not Giving a F*ck: A Counterintuitive Approach to Living a Good LifeVon EverandThe Subtle Art of Not Giving a F*ck: A Counterintuitive Approach to Living a Good LifeBewertung: 4 von 5 Sternen4/5 (5782)
- Grit: The Power of Passion and PerseveranceVon EverandGrit: The Power of Passion and PerseveranceBewertung: 4 von 5 Sternen4/5 (587)
- Hidden Figures: The American Dream and the Untold Story of the Black Women Mathematicians Who Helped Win the Space RaceVon EverandHidden Figures: The American Dream and the Untold Story of the Black Women Mathematicians Who Helped Win the Space RaceBewertung: 4 von 5 Sternen4/5 (890)
- The Yellow House: A Memoir (2019 National Book Award Winner)Von EverandThe Yellow House: A Memoir (2019 National Book Award Winner)Bewertung: 4 von 5 Sternen4/5 (98)
- On Fire: The (Burning) Case for a Green New DealVon EverandOn Fire: The (Burning) Case for a Green New DealBewertung: 4 von 5 Sternen4/5 (72)
- The Little Book of Hygge: Danish Secrets to Happy LivingVon EverandThe Little Book of Hygge: Danish Secrets to Happy LivingBewertung: 3.5 von 5 Sternen3.5/5 (399)
- A Heartbreaking Work Of Staggering Genius: A Memoir Based on a True StoryVon EverandA Heartbreaking Work Of Staggering Genius: A Memoir Based on a True StoryBewertung: 3.5 von 5 Sternen3.5/5 (231)
- Team of Rivals: The Political Genius of Abraham LincolnVon EverandTeam of Rivals: The Political Genius of Abraham LincolnBewertung: 4.5 von 5 Sternen4.5/5 (234)
- Devil in the Grove: Thurgood Marshall, the Groveland Boys, and the Dawn of a New AmericaVon EverandDevil in the Grove: Thurgood Marshall, the Groveland Boys, and the Dawn of a New AmericaBewertung: 4.5 von 5 Sternen4.5/5 (265)
- The Hard Thing About Hard Things: Building a Business When There Are No Easy AnswersVon EverandThe Hard Thing About Hard Things: Building a Business When There Are No Easy AnswersBewertung: 4.5 von 5 Sternen4.5/5 (344)
- The Emperor of All Maladies: A Biography of CancerVon EverandThe Emperor of All Maladies: A Biography of CancerBewertung: 4.5 von 5 Sternen4.5/5 (271)
- The World Is Flat 3.0: A Brief History of the Twenty-first CenturyVon EverandThe World Is Flat 3.0: A Brief History of the Twenty-first CenturyBewertung: 3.5 von 5 Sternen3.5/5 (2219)
- The Unwinding: An Inner History of the New AmericaVon EverandThe Unwinding: An Inner History of the New AmericaBewertung: 4 von 5 Sternen4/5 (45)
- The Gifts of Imperfection: Let Go of Who You Think You're Supposed to Be and Embrace Who You AreVon EverandThe Gifts of Imperfection: Let Go of Who You Think You're Supposed to Be and Embrace Who You AreBewertung: 4 von 5 Sternen4/5 (1090)
- The Sympathizer: A Novel (Pulitzer Prize for Fiction)Von EverandThe Sympathizer: A Novel (Pulitzer Prize for Fiction)Bewertung: 4.5 von 5 Sternen4.5/5 (119)
- Her Body and Other Parties: StoriesVon EverandHer Body and Other Parties: StoriesBewertung: 4 von 5 Sternen4/5 (821)
- Nizhal Tree Lists PDFDokument5 SeitenNizhal Tree Lists PDFMaheshChakravarthiNoch keine Bewertungen
- This Content Downloaded From 187.212.26.79 On Thu, 11 Aug 2022 02:01:26 UTCDokument62 SeitenThis Content Downloaded From 187.212.26.79 On Thu, 11 Aug 2022 02:01:26 UTCMAURICIO ARMANDO PAEZ SANTOSNoch keine Bewertungen
- Orthopaedics MCQSFDokument74 SeitenOrthopaedics MCQSFpt.mahmoudNoch keine Bewertungen
- Ch.2.2 Ecological Pyramid WorksheetDokument2 SeitenCh.2.2 Ecological Pyramid Worksheet무루파털뭉치Noch keine Bewertungen
- Quikbib - Pea Aphids 139Dokument11 SeitenQuikbib - Pea Aphids 139Rusdi HidayatNoch keine Bewertungen
- Oxidative Stress - Molecular Mechanisms and Biological EffectsDokument374 SeitenOxidative Stress - Molecular Mechanisms and Biological EffectsEML100% (2)
- Lab Assesment Number 2Dokument3 SeitenLab Assesment Number 2Aakash ReddyNoch keine Bewertungen
- NBME Samples Qs - PathologyDokument7 SeitenNBME Samples Qs - PathologyAli AlshehhiNoch keine Bewertungen
- Phytostabilization of Nickel by The Zinc and Cadmium Hyperaccumulator SolanumDokument7 SeitenPhytostabilization of Nickel by The Zinc and Cadmium Hyperaccumulator SolanumYogi PernandaNoch keine Bewertungen
- A Catalogue of All The Plants Part. 2Dokument623 SeitenA Catalogue of All The Plants Part. 2Abanti MitraNoch keine Bewertungen
- Animal Classification (Porifera-Echinodermata)Dokument164 SeitenAnimal Classification (Porifera-Echinodermata)9415697349Noch keine Bewertungen
- Astm A194Dokument3 SeitenAstm A194luizz100% (1)
- Enzymes Form 3Dokument4 SeitenEnzymes Form 3Ralph MuchingamiNoch keine Bewertungen
- Melanin Myth #3 Melanin Is CarbonDokument3 SeitenMelanin Myth #3 Melanin Is CarbonNnamdi Azikiwe0% (1)
- 1st Year DPT Course ContentDokument20 Seiten1st Year DPT Course ContentKhanNoch keine Bewertungen
- Molecular Drug Targets-1Dokument75 SeitenMolecular Drug Targets-1Phoebe Llamelo100% (1)
- Cell Structure FunctionDokument4 SeitenCell Structure FunctionLindley PareñoNoch keine Bewertungen
- Protein Function Prediction IntroductionDokument20 SeitenProtein Function Prediction IntroductionEndah Sekar PalupiNoch keine Bewertungen
- JurnalDokument7 SeitenJurnalDangdut ManiaNoch keine Bewertungen
- Patrick TB Ch08Dokument12 SeitenPatrick TB Ch08Marissa100% (1)
- As ISO 10993.5-2002 Biological Evaluation of Medical Devices Tests For in Vitro CytotoxicityDokument8 SeitenAs ISO 10993.5-2002 Biological Evaluation of Medical Devices Tests For in Vitro CytotoxicitySAI Global - APACNoch keine Bewertungen
- Rhode Island Coastal Buffer Zone Planting Guide - University of Rhode IslandDokument14 SeitenRhode Island Coastal Buffer Zone Planting Guide - University of Rhode IslandFree Rain Garden Manuals and MoreNoch keine Bewertungen
- Science9 q1 Mod4 SDOv2Dokument32 SeitenScience9 q1 Mod4 SDOv2Kaira Stella100% (1)
- Oncothermia: A New Paradigm in Cancer TherapiesDokument31 SeitenOncothermia: A New Paradigm in Cancer TherapiesCureus100% (1)
- Understanding BiodiversityDokument10 SeitenUnderstanding BiodiversityJeffrish raidnNoch keine Bewertungen
- Practice Test Questions Downloaded From FILIPINO NURSES CENTRALDokument26 SeitenPractice Test Questions Downloaded From FILIPINO NURSES CENTRALFilipino Nurses CentralNoch keine Bewertungen
- d20 Adamant Entertainment PosthumanDokument27 Seitend20 Adamant Entertainment Posthumanjoaam100% (3)
- Microorganisms are vital to humans and the environmentDokument29 SeitenMicroorganisms are vital to humans and the environmentNikita SinghNoch keine Bewertungen
- Gas Exchange Self StudyDokument5 SeitenGas Exchange Self Study4E-27 Tsoi Cheuk Ying (Ada)Noch keine Bewertungen
- Social Network Influence On YouthDokument3 SeitenSocial Network Influence On YouthKhairun NisaNoch keine Bewertungen