Beruflich Dokumente
Kultur Dokumente
Use of CCA and CA
Hochgeladen von
Sanet Janse van VuurenOriginalbeschreibung:
Copyright
Verfügbare Formate
Dieses Dokument teilen
Dokument teilen oder einbetten
Stufen Sie dieses Dokument als nützlich ein?
Sind diese Inhalte unangemessen?
Dieses Dokument meldenCopyright:
Verfügbare Formate
Use of CCA and CA
Hochgeladen von
Sanet Janse van VuurenCopyright:
Verfügbare Formate
Accepted for publication in Oecologia Pierre Legendre Eugene D.
Gallagher
Ecologically meaningful transformations for ordination of species data
Pierre Legendre (*) Dpartement de sciences biologiques, Universit de Montral, C.P. 6128, succursale Centre-ville, Montral, Qubec H3C 3J7, Canada e-mail: Pierre.Legendre@umontreal.ca Fax: 514-343-2293 Eugene D. Gallagher Department of Environmental, Coastal & Ocean Sciences, University of Massachusetts at Boston, Boston, Massachusetts 02125, USA Abstract This paper examines how to obtain species biplots in unconstrained or constrained ordination without resorting to the Euclidean distance (used in principal component analysis, PCA, and redundancy analysis, RDA) or the chi-square distance (preserved in correspondence analysis, CA, and canonical correspondence analysis, CCA) which are not always appropriate for the analysis of community composition data. To achieve this goal, transformations are proposed for species data tables. They allow ecologists to use ordination methods such as PCA and RDA, which are Euclidean-based, for the analysis of community data, while circumventing the problems associated with the Euclidean distance, and avoiding CA and CCA which present problems of their own in some cases. This allows the use of the original (transformed) species data in RDA carried out to test for relationships with explanatory variables (i.e. environmental variables, or factors of a multifactorial analysis-of-variance model); ecologists can then draw biplots displaying the relationships of the species to the explanatory variables. Another application is to allow the use of species data in other methods of multivariate data analysis which optimize a least-squares loss function; an example is K-means partitioning. Key words Biplot diagram Canonical correspondence analysis Correspondence analysis Principal component analysis Redundancy analysis
Transformations for species data
Introduction
Correspondence analysis (CA) and canonical correspondence analysis (CCA) are widely used to obtain unconstrained or constrained ordinations of species abundance data tables and the corresponding biplots or triplots which are extremely useful for ecological interpretation (Fig. 1a, c). Empirical work during the 1970s established that correspondence analysis was appropriate for such data, while ter Braak (1985) showed that the chi-square distance preserved in CA provided a good approximation for species with unimodal distributions along a single environmental gradient. There is a problem with this metric, however: a difference between abundance values for a common species contributes less to the distance than the same difference for a rare species, so that rare species may have an unduly large inuence on the analysis (Greig-Smith 1983, Legendre and Legendre 1998, ter Braak and Smilauer 1998). To avoid this, users of CA and CCA may remove the rarest species from the analysis, or resort to empirical methods giving small weights to rare species, as found for instance in the ordination program Canoco (ter Braak and Smilauer 1998). The chi-square distance is not unanimously accepted among ecologists: using simulated data, Faith et al. (1987) concluded that it was one of the worst distances for community composition data. Alternatives to CA and CCA are principal component analysis (PCA, for unconstrained ordination) and redundancy analysis (RDA, for constrained ordination) (Fig. 1a, c). In the fulldimensional space, these methods preserve the Euclidean distances among sites. For the analysis of sites representing short gradients, PCA and RDA may be suitable. For longer gradients, many species are replaced by others along the gradient and this generates lots of zeros in the species data table. Community ecologists have repeatedly argued that the Euclidean distance (and thus PCA and RDA) is inappropriate for raw species abundance data involving null abundances (e.g., Orlci 1978, Wolda 1981, Legendre and Legendre 1998; Table 1). For that reason, CCA is often the method favoured by researchers who are analysing compositional data, despite the problem posed by rare species. Other alternatives are available for unconstrained ordination analysis. One may compute a
Transformations for species data
resemblance matrix (similarity or distance) among sites using any of a number of resemblance coefcients that are appropriate for species presence-absence or abundance data; see Legendre and Legendre (1998) for a review. Following this, principal coordinate analysis (PCoA) or nonmetric multidimensional scaling (NMDS) can be used to obtain an ordination in a small number of dimensions, usually two or three. To obtain biplots of species and sites from PCoA or NMDS, one can (1) compute correlations between the original species vectors (i.e., the vectors whose i-th components are the count of a species at site i) and the site scores along the PCoA or NMDS ordination axes and scale these correlations as described in Eq. 14 (below); or (2) use the site scores along the two or three PCoA or NMDS ordination axes retained for ordination, together with the original species data, and carry out a PCA of this larger data table in which the original species will be treated as supplementary variables having weights of zero in the analysis; the use of supplementary variables in PCA is described, for instance, in ter Braak and Smilauer (1998) and Legendre and Legendre (1998). Resemblance matrices cannot be used directly in canonical ordination, however. Legendre and Anderson (1999) have proposed a solution to this problem, called distance-based redundancy analysis (db-RDA, Fig. 1e): (1) compute a matrix of distances Dij among sites using a measure appropriate to species data, e.g. the Steinhaus/Odum/Bray-Curtis measure (called Bray-Curtis for simplicity) which is often the preferred choice of ecologists; (2) compute all the principal coordinates, using PCoA; they preserve the original distances Dij in full ordination space; if negative eigenvalues and corresponding complex-number axes are produced during eigenvalue decomposition, a correction can be applied to the distance matrix to eliminate them; (3) use RDA to analyse the relationship between the principal coordinates, representing the species data, and the explanatory variables. db-RDA is well-suited to test the signicance of relationships between the explanatory and response data tables, but not to the production of biplots or triplots of species, sites and environmental variables, which may be needed for interpretation (Gabriel 1982, ter Braak 1994), because the species matrix is replaced in step 3 by another matrix whose columns are principal coordinates. Each column now represents a non-linear combination of the original
Transformations for species data
species, so that their roles cannot easily be untangled. The db-RDA approach can be used either for regular redundancy analysis of community composition against environmental variables, or to obtain a Manova-like analysis in which the factors of the Manova are coded in the matrices of environmental variables and covariables of the canonical analysis. See Legendre and Anderson (1999) for details. The present paper describes transformations of the species data that allow ecologists to use ordination methods such as PCA and RDA, which are Euclidean-based, with community composition data containing many zeros (long gradients). These transformations offer alternatives, for ordination analysis of community data, to CA and CCA, which are based upon the chi-square metric (Fig. 1b). They allow the use of the original (transformed) species data in RDA to test the relationships with explanatory variables (i.e. environmental variables or factors of a multifactorial analysis-of-variance model, Fig. 1d), as an alternative to db-RDA (Fig. 1e), thus allowing one to draw biplots displaying the relationships of the species to the explanatory variables. An additional application is to allow the use of community composition data in other methods of multivariate analysis which optimize a least-squares loss function. An example is Kmeans partitioning which separates the objects (e.g., sampling sites) into groups obtained by minimizing the sum of the squared Euclidean distances of the objects to the group centroids.
Transform species composition data to obtain targeted (dis)similarity coefcients
Some (dis)similarity measures commonly used by community ecologists can be obtained by rst modifying the species data, then computing the Euclidean distance among sites on the modied data set. We are proposing here to transform the species presence-absence or abundance data and use the transformed data in PCA, RDA or K-means partitioning. The net result is an analysis that will preserve the chosen distances among objects. Not all similarity coefcients that have been proposed to analyse community structure data can be obtained through such a transformation, however; see the Discussion. Consider a species abundance data table Y = [yij] of size (n p) with sites (rows)
Transformations for species data
i = {1 n} and species (columns) j = {1 p}; the row sums are noted yi+ and the column sums y+j ; the overall sum is y++ . We will dene transformations of the data Y Y' such that the Euclidean distance (eq. 4) among the rows of the transformed data table Y' is equal to some other distance computed among the rows of the original data table Y (Fig. 2). In the remainder of this paper, we will consider only distance coefcients; if required by the software, the corresponding similarities can be obtained by S(i,j) = 1 D(i,j) or S(i,j) = 1 D2(i,j), after ranging the distances to the interval [0, 1] if necessary. The transformations described below are precursors of distances that have all been described as appropriate for community composition data. They allow users to retain the identity of the individual species in PCA or RDA biplots. 1. Chord distance The chord distance, proposed by Orlci (1967) and Cavalli-Sforza and Edwards (1967), is the Euclidean distance computed after scaling the site vectors to length 1, i.e. dividing each value by the norm, or length, of the vector. After normalization, the Euclidean distance between two objects (sites) is equivalent to the length of a chord joining two points within a segment of a hypersphere of radius 1. The formula for the chord distance between sites x1 and x2 across the p species is thus: 2 p y2 j y1 j D chord ( x 1, x 2 ) = -------------------- -------------------- (1) p p j = 1 2 2 y1 j y2 j j=1 j=1 The chord distance may also be computed using the following formula found in several textbooks and papers (e.g., Orlci 1967): y1 j y2 j j=1 2 1 -------------------------------------- p p 2 2 y1 j y2 j
j=1 j=1 p
D chord ( x 1, x 2 ) =
(2)
Transformations for species data It is clear from eq. 1 that if the data [yij] are rst transformed into [ y' ij ] as follows: y ij y' ij = ------------------p
(3)
yij
2 j=1
then the Euclidean distance D Euclidean ( x' 1, x' 2 ) =
j=1
( y' 1 j y' 2 j )
(4)
between row vectors of transformed data is identical to the chord distance between the original row vectors of species abundances. The inner part of eq. 2 is actually the cosine of the angle () between the two site vectors, normalized or not; this is easily derived from the scalar product of two vectors: b c = (length of b) (length of c) cos . So the chord distance may be written as: D chord = 2 ( 1 cos ) (5)
This distance is maximum when the two sites have no species in common; the normalized site vectors may then be represented by points at 90 from each other on the circumference of a sector of a circle (for two species) or the surface of a segment of a hypersphere (for p species) and the distance between the two sites is 2 . Trueblood et al. (1994) used a form of equation 3 in their
PCA-H method, with yij being the probability of sampling species j in sample i with a random draw of m individuals from the sample. They called the Euclidean distances between these transformed row vectors CNESS, the chord normalized expected species shared. CNESS is a metric version of Grassle and Smiths (1976) NESS similarity index. Orlcis chord distance equals CNESS when m = 1. 2. Chi-square metric and chi-square distance The chi-square metric is often used for clustering or ordination of species abundance data. Although this measure has no upper limit, it produces distances smaller than 1 in most cases. The formula is:
Transformations for species data
metric
( x 1, x 2 ) =
j=1
-------j ------- ------- y+ y y
1+ 2+
1 y1 j
y2 j 2
(6)
The inner part is the Euclidean distance computed on relative abundances, weighted by the inverse of the column (species) sums y+j . If a species j is rare, its column sum y+j is small and this species contributes a great deal to the sum of squares. If the data [yij] are transformed into [ y' ij ] as follows: y ij y' ij = -----------------y i+ y + j (7)
then the Euclidean distance (eq. 4) between row vectors of transformed data is identical to the chisquare metric (eq. 6) between the original row vectors of species abundances. The chi-square distance is the chi-square metric multiplied by the square root of the sum of abundances in the data table, y ++ . This distance is particularly important in numerical ecology
because it is the distance preserved in CA and CCA. These two distances are treated together because they only differ by a multiplicative constant. The formula for the chi-square distance is: ( x 1, x 2 ) =
distance
j=1
1 y1 j y2 j 2 ------------------- ------- ------- = y + j y ++ y 1+ y 2+
y ++
j=1
1 y1 j y2 j 2 ------ ------- ------- y + j y 1+ y 2+
(8)
If the data [yij] are transformed into [ y' ij ] as follows: y' ij = y ij y ++ -----------------y i+ y + j (9)
a little algebra shows that the Euclidean distance (eq. 4) between row vectors of transformed data is identical to the chi-square distance (eq. 8) between the original row vectors of species abundances. This is not to say that a PCA of data transformed as in eq. 9, or a PCoA of a matrix of chi-square distances, will exactly reproduce a CA ordination; the rotation of the data point distribution in CA does not maximize inertia in the same way as in PCA or PCoA. The transformation proposed here (eq. 9) had been described by Chardy et al. (1976) and used by Pinel-Alloul et al. (1995) to prepare matrices of phytoplankton (87 taxa) and sh (18 taxa) data, prior to using them as matrices of explanatory variables in a CCA where the
Transformations for species data dependent matrix was a table of zooplankton abundances (54 taxa). 3. Distance between species proles
A variant of the chi-square metric can be obtained by removing the standardization by the inverse of y+j . The species data are simply transformed into proles of relative frequencies before computing Euclidean distances. This equation does not give extra weight to the rare species, the most abundant species contributing predominantly to the sum of squares. The formula is: D species profiles ( x 1, x 2 ) = ------- ------- y y
1+ 2+ p
y1 j
y2 j 2
(10)
j=1
This formula is constructed in the same way as Eq. 1; it is the Euclidean distance (Eq. 4) between species proles. If the data [yij] are rst transformed into [ y' ij ] as follows: y ij y' ij = -----y i+ (11)
then the Euclidean distance (eq. 4) between rows of transformed data (which are also called compositional data) is identical to the distance between species proles computed on the original species abundance data (eq. 10). 4. Hellinger distance The Hellinger distance is also a measure recommended for clustering or ordination of species abundance data (Rao 1995). The formula is: D Hellinger ( x 1, x 2 ) =
j=1
y1 j y2 j ------- ------y 1+ y 2+
(12)
Rao (1995) recommends it as a basis for a new ordination method. Using simulations, Legendre and Legendre (1998) concluded that for linear ordination, the Hellinger distance offers a better compromise between linearity and resolution than the chi-square metric and the chi-square distance. If the data [yij] are rst transformed into [ y' ij ] as follows:
Transformations for species data
y' ij =
y ij -----y i+
(13)
then the Euclidean distance (eq. 4) between row vectors of transformed data is identical to the Hellinger distance between the original row vectors of species abundances. The transformations described in this section form a set in the sense that the corresponding distances have all been recommended for analysis of community composition data and they can all be obtained by transforming the species abundance data followed by computation of Euclidean distances between the rows of transformed data. Other coefcients, described for instance in Legendre and Legendre (1998), that are appropriate for community composition analysis cannot be obtained using simple transformations of the species abundance data.
Example
To illustrate the differences among ordination methods, an articial ecological gradient was created by generating abundances for 9 species at 19 sites along a transect (Fig. 3a). Species 2 to 4, represented by 36 individuals each, replace each other along the gradient. Species 1 and 5 have the same kind of distribution and appear at the ends of the transect. Rare species 6 to 9 occur in narrow ranges of conditions along the gradient; they are represented by 2 to 5 individuals. Distance functions that are suitable for the analysis of community composition data should, minimally, be able to produce reasonable reconstructions of such a simple gradient. This can be assessed by looking either at the distance matrices themselves (Fig. 3) or at the biplots (Fig. 4). When examining distance matrices, one expects distances to increase monotonically as sites get further apart, until a maximum is reached for sites that have no species in common. This can be displayed using graphs of the computed ecological distances (ordinate) against the true geographic distances along the transect (abscissa). A model of the relationship can be drawn by joining the mean ecological distances computed for each geographic distance. If sampling has taken place on a geographic surface instead of a transect, the geographic distances will not fall into a small number of discrete distances; a smooth function can be drawn using moving
Transformations for species data
10
averages, splines, or LOWESS smoothing. We call this type of graph a diastemogram, from the Greek (diastema) distance, and (gramma) drawing. In Fig. 3, the diastemogram function is monotonically increasing for the chord, chi-square, Hellinger and Bray-Curtis distances, as sites get farther away along the gradient. Even though the Bray-Curtis distance cannot be obtained using one of the transformations of the previous section, it was included in Fig. 3 for comparison and reference because of its wide use in community ecology. The diastemogram function is not monotonic for the Euclidean distance and the distance between species proles. On the other hand, the coefcient of determination (R2) measures how much of the variance of the ecological distance matrix is explained by the diastemogram function; the value of R2 is low for the Euclidean distance and the distance between species proles. These are indications that these two distances should not be used to represent at least this particular ecological gradient. Note, however, that PCA based on species proles can be interpreted in terms of alpha and beta diversity (ter Braak 1983). In our example, R2 is the highest for the Hellinger distance, followed by the Bray-Curtis and chord distances; the chi-square distance, which does not reach an asymptote, comes last by that criterion. The diastemogram function is monotonic for these three distances. So, the best choices for this example seem to be the Hellinger and chord distances, for which ordinations can be obtained through the simple transformations described in the previous section followed by PCA or RDA (Fig. 1b and 1d), or the Bray-Curtis distance for which ordination diagrams can be obtained by PCoA; biplots of species and sites are more difcult to obtain, however, in that case. Another criterion for the comparison of distance measures is the importance given to rare species in the analysis. If rare species are well-sampled and truly rare, they may be used as indicators of the conditions that may exist at some sites only. In that case, they should receive high weight, as they do in CA and, to a lesser extent, in PCA after the chi-square transformation (Table 2). On the contrary, when rare species are observed sporadically at some sites, but, as the result of sampling error, not at others where they are also present, it is unwise to give them high weight in the analysis. This phenomenon is exacerbated in environments where sampling is
Transformations for species data
11
conducted blindly for instance, in aquatic and soil ecology. Coefcients such as the Euclidean, chord, species prole or Hellinger distances do not give high weights to the rare species (Table 2). The combined effect of the various types of transformations and the most commonly used ordination methods was assessed by carrying out ordination of the data, rst by PCA and CA, then by PCA after transforming the data as described above. We observe the following: After PCA of the raw data (Fig. 4a), axis 1 displays the gradient with strong inward folding of the sites at the ends of the gradient (horseshoe effect). Because the Euclidean distance function considers double zeros as an indication of similarity, PCA brings together sites from the two ends of the gradient that have no species in common. PCA after the transformation into species proles (Eq. 11) produces similar results (Fig. 4e). The inadequacy of ordinations displaying strong horseshoes has been discussed in the ecological literature since Goodall (1954). CA displays the gradient correctly along axis 1 (Fig. 4b); so does PCA on data transformed by Eq. 9, where the ordination preserves the chi-square distance among sites (Fig. 4c); Eq. 7 leads to similar results (not shown). The small differences in the ordination of sites between Figs. 4b and 4c is due to the fact that CA does not dene inertia in the same way and, so, does not apportion it among axes in the same way as PCA. PCAs on data transformed using Eqs. 3 (chord transformation, Fig. 4d) and 13 (Hellinger transformation, Fig. 4f) produce good representations of the gradient along axis 1, with some inward folding of three sites at both ends of the transect. PCoA of a Bray-Curtis distance matrix is shown in Fig. 4g for comparison. Linear correlations were computed between the original species vectors and the rst two principal coordinates. The correlations were weighted as follows to obtain the species scores for the biplot: SpeciesScorejk = rjksj /sk (14)
where rjk is the correlation between species j and site score vector k, sj is the standard deviation of species j, and sk is the standard deviation of site score vector k. The term sk adjusts the species scores to the scaling used in any particular analysis: the variance of site score vector k is k in PCoA and in PCA distance biplots (k = eigenvalue), whereas it is 1 in PCA correlation biplots.
Transformations for species data
12
For the present example, the ordinations of sites obtained for the chord and Hellinger transformations have inward folding of three sites at both ends of the transect (horseshoe effect); the horseshoe is not stronger than in the ordination obtained using the Bray-Curtis distance (Fig. 4). The horseshoe effect is much stronger in the case of the Euclidean distance and the species prole transformation. CA as well as PCA after chi-square distance transformation produced no horseshoe effect in this example. Noticeably, the fraction of variance of the species data accounted for by the rst two ordination axes is much lower in CA and in the PCA following chisquare distance transformation (49-50%) than in the other PCAs (67-71%). The Bray-Curtis PCoA ordination is intermediate in that respect (57%). In the biplots, where the rst two axes only were used, all methods based upon PCA gave a fair representation of the relative numerical importance of the rare species. This included the PCA after chi-square distance transformation of the data: even though the rare species have high weights in the analysis (higher variance in Table 2 than in the other PCA results), they only load heavily on the lesser ordination axes. In CA, where the rare species have high weights (Table 2), the importance of a species for an ordination subspace is given, e.g. in the program Canoco, by the Cumulative t per species as fraction of variance of species. Here again, the rare species only load heavily on the last ordination axes. Fig. 4b (CA) shows the species at the centroids of the sites where they are present; compare Figs. 3a and 4b. The lengths of the lines joining the species to the origin is not a measure of their importance in the analysis; in CA biplots, species scores are slopes with respect to the axes (ter Braak and Smilauer 1998, Eq. 6.17).
Discussion
The transformations described above are precursors of distances that have all been described as appropriate for community composition data. This is not to say that these ve are the only appropriate distances for species abundance data; see for instance Faith et al. (1987) or Legendre and Legendre (1998) for reviews of appropriate coefcients. These distances, however, can be obtained through transformations that allow users to retain the identity of the individual species in
Transformations for species data
13
biplots. Prior to computing these transformations, any of the standardizations investigated by Faith et al. (1987) may also be used. Theoretical considerations The distance functions that can be obtained by transformation of the species data followed by calculation of Euclidean distances have some interesting mathematical properties: 1) All distance functions pertaining to this group are Euclidean, meaning that the distances among objects that they produce can be entirely represented in Euclidean space. A representation is Euclidean when principal coordinate analysis of the distance matrix does not produce negative eigenvalues. These concepts are explained by Legendre and Legendre (1998), for instance. 2) The distances corresponding to the transformations described above can be computed on presence-absence species data. Hence the transformations can also be applied to this type of data. 3) Coefcients that are non-Euclidean cannot be obtained through transformation of the data followed by calculation of Euclidean distances. If it were possible to do so, the resulting distances would be Euclidean, while they are not. In particular, distances which are one-complements of similarity coefcients for binary (0-1) data, such as the simple matching, Jaccards, or Srensens coefcients, cannot be obtained through such transformations since none of them are Euclidean (Gower and Legendre 1986; Legendre and Legendre 1998, Table 7.2); Srensens coefcient is not even a metric. The widely-used Bray-Curtis coefcient for community data, which ranked among the best of the coefcients studied by Faith et al. (1987), cannot be obtained through such a transformation because it is neither Euclidean nor metric; likewise for the NESS similarity coefcient of Grassle and Smith (1976). To use these otherwise excellent coefcients in constrained ordination, one should use either the db-RDA approach of Legendre and Anderson (1999) or the alternative computation procedure proposed by McArdle and Anderson (2001). 4) The coefcients that can be obtained by data transformation are those that can be expressed as an equation where each value yij is weighted by some function of the values in the same row and/or column (e.g., the sum, or the sum of squares, of the values). The sum of all values y++ may
Transformations for species data
14
also be included in the transformation. As a result, these coefcients only compare species proles, normalized in various ways. To obtain distances that preserve total abundances at the sites, instead of normalized proles, one has to use the coefcient of Bray-Curtis or that of Kulczynski; see Faith et al. (1987) or Legendre and Legendre (1998) for details. Practical considerations For the analysis of community gradients, it does not matter that an analysis gives high weights to the rare species when the end-product is simply a reduced-space ordination diagram. In CA or CCA, and in PCA or RDA based upon the chi-square metric (Eq. 7) or the chi-square distance transformation (Eq. 9), the rarest species are well tted by the axes with the smallest eigenvalues. The contributions of these species to the rst few axes, used for reduced-space ordination, are small. The weights given to rare species do matter, however, when the end-product is a test of signicance of the relationship between species composition and a set of explanatory variables, or a test of factors in a multiple analysis-of-variance model following the db-RDA approach of Legendre and Anderson (1999). CCA, or the chi-square distance transformation followed by RDA, should not be used unless one specically wants to give high weight to the rare species that may indicate the presence of particular environmental conditions. The chord and Hellinger transformations are appropriate alternatives giving low weights to rare species. In our example, the diastemogram functions for these two transformations showed that the resulting distances were monotonically related to the geographic distances along the gradient, as in the case of the Bray-Curtis coefcient; they reached an asymptote for sites that had no species in common; they produced little horseshoe effect in ordinations; and they allowed the representation of species and sites in biplots. For simple ordination analysis, the difference between CA and PCA after the chi-square distance transformation is small. With appropriate scalings of the species and site scores, the same distance is preserved in the two forms of analysis, but the eigenvalues may differ slightly. There is a more important difference in canonical ordination, however: the multiple regressions of the
Transformations for species data
15
species on the set of explanatory variables are done with weights in CCA; this is not the case in RDA. The weights in CCA are given by a diagonal matrix containing the square roots of the row sums of the species data table. This means that a site where many individuals have been observed contributes more to the regression than a site with few individuals. CCA should only be used when the sites have approximately the same number of individuals, or when one explicitly wants to give high weight to the richest sites. This problem of CCA was one of our incentives for looking for alternative methods for canonical ordination of community composition data. RDA based upon the transformations proposed in this paper (including the chi-square metric or distance transformations) offers an alternative solution. PCA of raw data, which preserves the Euclidean distance among sites in full-dimensional space, is inappropriate for species composition data. PCA of the species proles allows a quick view of both alpha diversity (as Simpsons diversity) and beta diversity (at the cost of severe distortion at the ends of long gradients). Applications Here are some cases where the proposed data transformations may be useful for the analysis of species abundance tables, or other types of frequency data: In unconstrained or canonical ordination, when one does not wish to use the chi-square distance preserved by CA and CCA because of the differential weighting of rare species. In canonical ordination, when one does not wish to use the CCA weighting system for sites. When RDA or CCA is used to depict biotic control through top-down or bottom-up interactions (Lindeman 1942, Southwood 1987). In such studies, the analysis of a species matrix Y is constrained using another species matrix X representing predators or prey. The species data in X need to be transformed in such a way that the Euclidean distances among rows of X correspond to meaningful distances in species space, before X is used in a linear model; the transformations proposed in this paper can be used to do this. See Pinel-Alloul et al. (1995) for an example. The transformations described in this paper offer new ways of partitioning sites described by
Transformations for species data
16
species abundance data, i.e., dividing them into groups. One commonly used partitioning method is K-means, which is a Euclidean method minimizing a least-squares loss function. To preserve a distance function which is appropriate for community composition data, instead of the Euclidean distance which is inappropriate (Table 1), one can, prior to K-means, transform the species abundances using equations 3, 7, 9, 11 or 13. Theoretical criteria are not known at the moment that would allow one to select the best distance function (or data transformation) for any specic situation. In canonical analysis, one may empirically select the transformation which leads to the highest fraction of explained variation. Computer programs to carry out these transformations are available on the WWWeb sites <http://www.fas.umontreal.ca/biol/legendre/> (Fortran source code and compiled versions) and <http://www.es.umb.edu/edgwebp.htm> (Matlab code). Acknowledgements Thanks to Elaine Hooper who motivated this investigation by asking how to represent the original species in a biplot after db-RDA; to David W. Roberts who proposed improvements to the example data set used in this paper and to the ways of showing differences among distance functions; to Philippe Casgrain who incorporated diastemograms into The R Package; and to Daniel Borcard and Mark Burgman for comments on the manuscript. This research was supported by NSERC grant number OGP7738 to P. Legendre.
References
Cavalli-Sforza LL, Edwards AWF (1967) Phylogenetic analysis: models and estimation procedures. Evolution 21:550-570 Chardy P, Glemarec M, Laurec A (1976) Application of inertia methods to benthic marine ecology: practical implications of the basic options. Estuarine Coastal Shelf Sci. 4:179-205. Faith DP, Minchin PR, Belbin L (1987) Compositional dissimilarity as a robust measure of ecological distance. Vegetatio 69:57-68 Gabriel KR (1982) Biplot. In: Kotz S, Johnson NL (eds) Encyclopedia of statistical sciences, Vol. 1. Wiley, New York, pp 263-271
Transformations for species data
17
Goodall DW (1954) Objective methods for the classication of vegetation. III. An essay in the use of factor analysis. Aust. J. Bot. 2:304-324 Gower JC, Legendre P (1986) Metric and Euclidean properties of dissimilarity coefcients. J. Classif. 3:5-48 Grassle JF, Smith W (1976) A similarity measure sensitive to the contribution of rare species and its use in investigation of variation in marine benthic communities. Oecologia 25:13-25 Greig-Smith P (1983) Quantitative plant ecology. Third edition. Blackwell, London Legendre P, Anderson MJ (1999) Distance-based redundancy analysis: testing multispecies responses in multifactorial ecological experiments. Ecol. Monogr. 69:1-24 Legendre P, Legendre L (1998) Numerical ecology. Second English edition. Elsevier, Amsterdam Lindeman RL (1942) The trophic-dynamic aspect of ecology. Ecology 23:399-418 McArdle BH, Anderson MJ (2001) Fitting multivariate models to community data: a comment on distance-based redundancy analysis. Ecology 82:290-297 Orlci L (1967) An agglomerative method for classication of plant communities. J. Ecol. 55:193-205 Pinel-Alloul B, Niyonsenga T, Legendre P (1995) Spatial and environmental components of freshwater zooplankton structure. coscience 2:1-19 Rao CR (1995) A review of canonical coordinates and an alternative to correspondence analysis using Hellinger distance. Qestii (Quaderns dEstadstica i Investigaci Operativa) 19:23-63 Southwood TRE (1987) The concept and nature of the community. In: Gee JHR, Giller PS (eds) Organization of communities: past and present. Blackwell, Oxford, pp 3-27 ter Braak CJF (1983) Principal components biplots and alpha and beta diversity. Ecology 64:454462. ter Braak CJF (1985) Correspondence analysis of incidence and abundance data: properties in terms of a unimodal response model. Biometrics 41:859-873 ter Braak CJF (1994) Canonical community ordination. Part I: Basic theory and linear methods. coscience 1:127-140
Transformations for species data
18
ter Braak CJF, Smilauer P (1998) CANOCO reference manual and users guide to Canoco for Windows Software for canonical community ordination (version 4). Microcomputer Power, Ithaca, New York Trueblood DD, Gallagher ED, Gould DM (1994) Three stages of seasonal succession on the Savin Hill Cove mudat, Boston Harbor. Limnol. Oceanogr. 39:1440-1454 Wolda H (1981) Similarity indices, sample size and diversity. Oecologia 50:296-302
Transformations for species data
19
Table 1 Species abundance paradox, modied from Orlci (1978). The paradox is that the Euclidean distance between sites 1 and 2, which have no species in common, is smaller than that between sites 1 and 3 which share species 2 and 3. This example shows that the Euclidean distance is not appropriate for species community composition data containing zeros. With the other coefcients used here, the distance between sites 1 and 2 is larger than between sites 1 and 3, and the distance between sites 1 and 2 is the same as between sites 2 and 3, or very nearly so.
Species abundance paradox data (3 sites, 3 species)
Site 1 Site 2 Site 3
Species 1 0 1 0
Species 2 1 0 4
Species 3 1 0 8
Distance function D Euclidean D chord D D
metric distance
2 2
D(Site 1, Site 2) 1.7321 1.4142 1.0382 4.0208 1.2247 1.4142
D(Site 1, Site 3) 7.6158 0.3204 0.0930 0.3600 0.2357 0.1697
D(Site 2, Site 3) 9.0000 1.4142 1.0352 4.0092 1.2472 1.4142
D species profiles D Hellinger
Transformations for species data
20
Table 2 Fraction of the total variance occupied by each species, either for the raw species data from Fig. 3a (in the case of PCA and CA), or after the stated transformation (tr.) of the data. Each row sums to 1. The variance of a species vector measures its relative importance in the analysis. Species 6 to 9 are the rare species.
Sp. 1 PCA (original data) CA (original data)1 Chord tr. (Eq. 3) Chi-square tr. (Eq. 9) Prole tr. (Eq. 11) Hellinger tr. (Eq. 13)
Sp. 2
Sp. 3
Sp. 4
Sp. 5
Sp. 6
Sp. 7
Sp. 8
Sp. 9
0.1070 0.2434 0.2434 0.2434 0.1070 0.0032 0.0081 0.0164 0.0281 0.1060 0.0355 0.0349 0.0344 0.1037 0.1725 0.1725 0.1725 0.1679 0.1248 0.2303 0.2245 0.2192 0.1208 0.0065 0.0148 0.0249 0.0343 0.1686 0.1410 0.1393 0.1382 0.1644 0.0428 0.0584 0.0700 0.0773 0.1143 0.2458 0.2427 0.2409 0.1114 0.0041 0.0085 0.0136 0.0187 0.1216 0.2161 0.2126 0.2100 0.1166 0.0206 0.0284 0.0346 0.0395
ter Braak and Smilauer (1998, Eq. 6.36).
Transformations for species data
21
Unconstrained ordination of species data
(a) Classical approach Ordination biplot
Y = Raw data (sites x species)
Short gradients: CA or PCA Long gradients: CA
(b) Transformed data approach
Raw data (sites x species) Y=Transformed data (sites x species)
PCA
Representation of elements: Species = arrows Sites = symbols
Constrained ordination of species data
(c) Classical approach
X= Explanatory variables
Y = Raw data (sites x species)
Short gradients: CCA or RDA Long gradients: CCA Canonical ordination triplot
(d) Transformed data approach
Raw data (sites x species) Y=Transformed data (sites x species) X= Explanatory variables
RDA
(e) Distance-based RDA (db-RDA) approach
Raw data (sites x species) Distance matrix Representation of elements: Species = arrows Sites = symbols Explanatory variables = lines
PCoA
Y= (sites x principal coord.) X= Explanatory variables
RDA
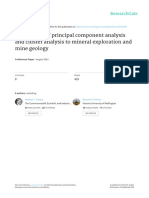
Fig. 1 Schematic comparison of techniques that can be used to obtain unconstrained (a, b) or constrained (c, d, e) ordination biplots or triplots of species data tables.
Transformations for species data
22
Chord (sites x species) transformation
Raw data
Transformed data (sites x species)
Chord distance among sites Chord distance matrix
Euclidean distance among sites
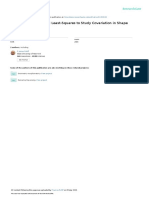
Fig. 2 Illustration of the role of the data transformations as a way of obtaining a given distance function. The example uses the chord distance.
Transformations for species data
23
Species abundance
8 6 4 2 0 1 Sp. 6 3 5 Sp. 7 7 Sp. 8 9 11 Sampling sites 13 15 Sp. 9 17 Species 1 Species 2 Species 3 Species 4
(a)
Species 5
19
Euclidean distance
12.0 8.0 6.0 4.0 2.0 0.0 0 2 4 6 8 10 12 14 16 True geographic distance 18 20 Chord distance 10.0
(b) R2 = 0.637
1.6 1.4 1.2 1.0 0.8 0.6 0.4 0.2 0.0 0 2 4
(c) R2 = 0.827
6 8 10 12 14 16 True geographic distance 18 20
4.0 3.0 2.0 1.0 0.0 0 2 4 6 8 10 12 14 16 True geographic distance 18 20
(d) R2 = 0.656
1.6 1.4 1.2 1.0 0.8 0.6 0.4 0.2 0.0 0 2 4
Dist. between profiles
Chi-square distance
(e) R2 = 0.669
6 8 10 12 14 16 True geographic distance 18 20
1.6 1.4 1.2 1.0 0.8 0.6 0.4 0.2 0.0 0 2 4
Bray-Curtis distance
1.2 1.0 0.8 0.6 0.4 0.2 0.0 0 2 4 6 8 10 12 14 16 True geographic distance 18 20
Hellinger distance
(f) R2 = 0.954
6 8 10 12 14 16 True geographic distance 18 20
(g) R2 = 0.855
Fig. 3 Analysis of articial gradient data. (a) The gradient comprises 19 sites (numbers along abscissa) and 9 species (different symbols). (b-g) Diastemograms comparing true geographic distances (abscissa) to the computed ecological distances among sites (ordinate). The construction and interpretation of these graphs is described in the text.
Transformations for species data
3.50 2.00 2.00 10 9 1.00 11 7 8 13 6 0.00 5 4 3 Sp.1 2 Sp.4 2.00 2.00 1 Sp.5 17 18 19 1.00 1 11 Sp.3 12 2.25 9 10 1.00 Sp.7 Sp.8 7 8 Sp.3 13 Sp.7 Sp.8 Sp.2 Sp.6 Sp.1 Sp.9 Sp.4 14 15 16
24
Sp.3
(a)
(b)
10 9 11 12
(c)
1.00
8 Sp.8 7 Sp.7 Sp.6 6 3 Sp.2 5 4
12
0.00 5 4 13 1.00 14 17 16 15 3
Sp.2
Sp.4 Sp.9
14 15 16
Sp.6
Sp.5
0.25
Sp.9
17 2 18 19
Sp.1 Sp.5 21 19 18
PCA, raw data
0.00
CA, raw data
1.00 0.00 1.00 2.00 2.00 2.00
PCA, chi-square distance transformation
1.00 0.00 1.00 2.00
1.50 2.50
1.25
1.25
2.50
2.00 10 9 8 1.00 Sp.3 12 11
2.50
2.00 10 9 11 8 1.00 Sp.3 8 7 12 Sp.7 Sp.6 Sp.2 13 4 Sp.4 14 3 4 2 1 19 18 17 16 15 1.00 3 17 2 1 19 18 5 Sp.1 Sp.8 Sp.9 Sp.5
(d)
1.50
(e)
10 9 Sp.3 12 13 11
(f)
7 0.00 6 5 4 1.00 3 2 1 Sp.2
Sp.7 Sp.8 Sp.6 Sp.9 Sp.1 Sp.5 Sp.4
13 0.50 14 15 19 16 18 17 0.50 5 6 Sp.2 7 Sp.7 Sp.8 Sp.6 Sp.9 Sp.1 Sp.5
6 0.00
14 Sp.4 15 16
PCA, Chord distance transformation
2.00 2.00 1.00 0.00 1.00 2.00 1.50 2.00
PCA, species profile transformation
1.00 0.00 1.00 2.00 2.00 2.00
PCA, Hellinger distance transformation
1.00 0.00 1.00 2.00
0.75
10 9 0.38 8 Sp.3 11 12
(g)
7 0.00 6 5 4 3 0.38 2 1 19 Sp.7 Sp.8 Sp.6 Sp.9 Sp.2 Sp.1 Sp.5 Sp.4
13
Editor: The Figure caption could be fitted here.
15 16
14
18 17
PCoA, Bray-Curtis distance
0.75 0.75 0.38 0.00 0.38 0.75
Fig. 4 Ordination of articial gradient data from Fig. 3a. (a) PCA, correlation biplot. (b) CA, species at the centroids of the sites in the biplot (CA scaling type 2). (c-f) PCA of transformed data (as specied in each panel), correlation biplots; these biplots preserve the correlations among species but may expand or otherwise distort the distances among sites. (g) PCoA of a Bray-Curtis distance matrix with superimposed species scores; see text, Eq. 14. Dots, numbered 1-19, represent the sampling sites; they are connected to materialize the gradient. Species are represented by lines. (d-f) The lengths of the species lines have been multiplied by 3 for clarity. (g)The lengths of the species lines have been divided by 25. Percentage of variance in the data explained by ordination axes I and II: (a) 69%, (b) 49%, (c) 50%, (d) 67%, (e) 69%, (f) 71%, (g) 59%.
Das könnte Ihnen auch gefallen
- Ecologically Meaningful Transformations For Ordination of Species DataDokument11 SeitenEcologically Meaningful Transformations For Ordination of Species DataEldo SantosNoch keine Bewertungen
- Legendre-Gallagher2001 Article EcologicallyMeaningfulTransfor PDFDokument10 SeitenLegendre-Gallagher2001 Article EcologicallyMeaningfulTransfor PDFAna Carolina Martins PereiraNoch keine Bewertungen
- Oblig Critical Article PDFDokument22 SeitenOblig Critical Article PDFbrrntosNoch keine Bewertungen
- Measuring Abundance: Methods for the Estimation of Population Size and Species RichnessVon EverandMeasuring Abundance: Methods for the Estimation of Population Size and Species RichnessNoch keine Bewertungen
- CART Algorithm For Spatial Data - Application To Environmental and Ecological DataDokument12 SeitenCART Algorithm For Spatial Data - Application To Environmental and Ecological Datathomas.romaryNoch keine Bewertungen
- Stochastic Methods for Flow in Porous Media: Coping with UncertaintiesVon EverandStochastic Methods for Flow in Porous Media: Coping with UncertaintiesNoch keine Bewertungen
- Jackson 1993 - Stopping Rules in PCADokument12 SeitenJackson 1993 - Stopping Rules in PCAvojarufosiNoch keine Bewertungen
- Impact of Sample Size On Principal Component AnalyDokument18 SeitenImpact of Sample Size On Principal Component AnalySunkari ArikaNoch keine Bewertungen
- Ordination AnalysisDokument8 SeitenOrdination AnalysisCsaba EötvösNoch keine Bewertungen
- Gazley Et Al. (2015) - Application of Principal Component Analysis and Cluster Analysis To Mineral Exploration and Mine GeologyDokument10 SeitenGazley Et Al. (2015) - Application of Principal Component Analysis and Cluster Analysis To Mineral Exploration and Mine GeologygeodennysNoch keine Bewertungen
- Thesis 5 p60-72 CCA New MethodDokument14 SeitenThesis 5 p60-72 CCA New MethodLIDIA PUGLIESSANoch keine Bewertungen
- Bayesian Estimation of Recent Migration Rates After A Spatial ExpansionDokument10 SeitenBayesian Estimation of Recent Migration Rates After A Spatial ExpansionAlessandra MarinsNoch keine Bewertungen
- Modelling in GisDokument34 SeitenModelling in GisMadhu ChaudharyNoch keine Bewertungen
- Graphical Displays For Meta-AnalysisDokument15 SeitenGraphical Displays For Meta-Analysisdinh son myNoch keine Bewertungen
- Permanova PaperDokument15 SeitenPermanova PaperjpallotNoch keine Bewertungen
- ANDERSON. 2001. A New Method For Non-Parametric Multivariate Analysis of VarianceDokument23 SeitenANDERSON. 2001. A New Method For Non-Parametric Multivariate Analysis of VarianceIchtus Soluções AmbientaisNoch keine Bewertungen
- Ga 12526Dokument154 SeitenGa 12526Edgar LongNoch keine Bewertungen
- Carrolletal2011 PDFDokument10 SeitenCarrolletal2011 PDFElva Domínguez TorresNoch keine Bewertungen
- HCPC Husson JosseDokument17 SeitenHCPC Husson JosseZain AamirNoch keine Bewertungen
- Anderson MJ 2001.permanovaDokument23 SeitenAnderson MJ 2001.permanovaSuzana DinizNoch keine Bewertungen
- A Maximum Entropy Approach To Species Distribution Modeling: Research, 180 Park Avenue, Florham Park, NJ 07932Dokument8 SeitenA Maximum Entropy Approach To Species Distribution Modeling: Research, 180 Park Avenue, Florham Park, NJ 07932Alejandra AranaNoch keine Bewertungen
- Legendre Fortin MER 2010Dokument14 SeitenLegendre Fortin MER 2010Catalina Verdugo ArriagadaNoch keine Bewertungen
- Section 7.7 Transformation HellingerDokument7 SeitenSection 7.7 Transformation HellingerMichelle Vanessa Soto AvendañoNoch keine Bewertungen
- jaescre,+RAET 45 4066Dokument14 Seitenjaescre,+RAET 45 4066informacion.2023.11Noch keine Bewertungen
- Detrended Correspondence Analysis (Dca) : Steven M. HollandDokument14 SeitenDetrended Correspondence Analysis (Dca) : Steven M. HollandPaulaNoch keine Bewertungen
- Specification and Testing of Some Modified Count Data ModelsDokument25 SeitenSpecification and Testing of Some Modified Count Data ModelsSuci IsmadyaNoch keine Bewertungen
- Moline 1995Dokument20 SeitenMoline 1995CLELIA NARANJO FREIRENoch keine Bewertungen
- Ecography - 2007 - F Dormann - Methods To Account For Spatial Autocorrelation in The Analysis of Species DistributionalDokument20 SeitenEcography - 2007 - F Dormann - Methods To Account For Spatial Autocorrelation in The Analysis of Species DistributionalNicolas RuizNoch keine Bewertungen
- Simple Metric Sedimentary Statistics Used To Recognize Different EnvironmentsDokument21 SeitenSimple Metric Sedimentary Statistics Used To Recognize Different EnvironmentsMarcia S. S. CarrilloNoch keine Bewertungen
- Ecography - 2012 - Crase - A New Method For Dealing With Residual Spatial Autocorrelation in Species Distribution ModelsDokument10 SeitenEcography - 2012 - Crase - A New Method For Dealing With Residual Spatial Autocorrelation in Species Distribution ModelsZihao AnNoch keine Bewertungen
- Eftekhari 1-7Dokument7 SeitenEftekhari 1-7Umesh MogleNoch keine Bewertungen
- Journal of Petroleum Science and Engineering: Hong Tang, Christopher D. WhiteDokument6 SeitenJournal of Petroleum Science and Engineering: Hong Tang, Christopher D. WhiteWaleed EjazNoch keine Bewertungen
- Clarke & Ainsworth. 1993. A Method of Linking Multivariate Community Structure To Environmental Variables.Dokument15 SeitenClarke & Ainsworth. 1993. A Method of Linking Multivariate Community Structure To Environmental Variables.mariaNoch keine Bewertungen
- Functional Models For Regression Tree Leaves: Luís TorgoDokument9 SeitenFunctional Models For Regression Tree Leaves: Luís TorgoLoh Jia SinNoch keine Bewertungen
- Baker & King 2010Dokument13 SeitenBaker & King 2010Cíntia RibeiroNoch keine Bewertungen
- GOVOROV M. (1), GIENKO G. (2) (1) Vancouver Island University, NANAIMO, CANADA (2) University of Alaska Anchorage, Anchorage, United StatesDokument13 SeitenGOVOROV M. (1), GIENKO G. (2) (1) Vancouver Island University, NANAIMO, CANADA (2) University of Alaska Anchorage, Anchorage, United StatesradocajdorijanNoch keine Bewertungen
- Do Not Log-Transform Count DataDokument5 SeitenDo Not Log-Transform Count DataarrudajefersonNoch keine Bewertungen
- Vogt Et - Al. Functional Connectivity (2009)Dokument16 SeitenVogt Et - Al. Functional Connectivity (2009)Paloma EntepolaNoch keine Bewertungen
- Decades of Miscomputation in Genomic Clades and DistancesDokument15 SeitenDecades of Miscomputation in Genomic Clades and DistancesAnonymous lVQ83F8mCNoch keine Bewertungen
- Multidimensional Data AnalysisDokument24 SeitenMultidimensional Data AnalysisNovia WidyaNoch keine Bewertungen
- Application of Correspondence Analysis in The Assessment of Groundwater ChemistryDokument50 SeitenApplication of Correspondence Analysis in The Assessment of Groundwater ChemistrysagawarnaNoch keine Bewertungen
- DCA TutorialDokument10 SeitenDCA TutorialdeltochilumNoch keine Bewertungen
- 9Dokument9 Seiten9Mariana SantanaNoch keine Bewertungen
- Modeling of Species Distributions With Maxent: New Extensions and A Comprehensive EvaluationDokument15 SeitenModeling of Species Distributions With Maxent: New Extensions and A Comprehensive EvaluationDavid AarónNoch keine Bewertungen
- Hypothesis Tests For Principal Component Analysis When Variables Are StandardizedDokument20 SeitenHypothesis Tests For Principal Component Analysis When Variables Are StandardizedNikhilNoch keine Bewertungen
- XXX Report Version Oct 2012Dokument38 SeitenXXX Report Version Oct 2012zarcone7Noch keine Bewertungen
- 2blockPLS RohlfCortiDokument16 Seiten2blockPLS RohlfCortiattilal_5Noch keine Bewertungen
- A Comparison of Methods For The Statistical Analys PDFDokument24 SeitenA Comparison of Methods For The Statistical Analys PDFRiza Wara ElzatiNoch keine Bewertungen
- Stochastic and Deterministic Seismic Inversion Met PDFDokument10 SeitenStochastic and Deterministic Seismic Inversion Met PDFDaniel Mauricio Rojas CaroNoch keine Bewertungen
- Revell 2009.evol - EarlyviewDokument11 SeitenRevell 2009.evol - EarlyviewOzan KorkmazNoch keine Bewertungen
- Lab #4 Quantifying Spatial Pattern I ObjectivesDokument13 SeitenLab #4 Quantifying Spatial Pattern I Objectivessherko jaafariNoch keine Bewertungen
- (McRae Et Al, 2008) Using Circuit Theory To Model Connectivity in Ecology, Evolution, and ConservationDokument14 Seiten(McRae Et Al, 2008) Using Circuit Theory To Model Connectivity in Ecology, Evolution, and ConservationOscar Leonardo Aaron Arizpe VicencioNoch keine Bewertungen
- The Impact of Mathematics On Cellular and Molecular BiologyDokument10 SeitenThe Impact of Mathematics On Cellular and Molecular Biologydevasena1991Noch keine Bewertungen
- The Others in The Cluster But With Differences Between ClustersDokument5 SeitenThe Others in The Cluster But With Differences Between ClustersParth Hemant PurandareNoch keine Bewertungen
- Some Methods For Analyzing and Correcting For Spatial AutocorrelationDokument26 SeitenSome Methods For Analyzing and Correcting For Spatial AutocorrelationFrancisco Parada SrepelNoch keine Bewertungen
- 5d30 PDFDokument12 Seiten5d30 PDFivasitonNoch keine Bewertungen
- Mom MF-1Dokument79 SeitenMom MF-1AnbuNoch keine Bewertungen
- Journal of Building Information Modeling - Fall 2010Dokument40 SeitenJournal of Building Information Modeling - Fall 2010bimpireNoch keine Bewertungen
- Lesson Plan GagneDokument2 SeitenLesson Plan Gagnechiew gin siongNoch keine Bewertungen
- GEK 63383d Turbine Generator FoundationDokument61 SeitenGEK 63383d Turbine Generator Foundationbong2rm100% (6)
- Human BehaviorDokument14 SeitenHuman BehaviorJay Ann AguilarNoch keine Bewertungen
- PIC16F1946Dokument448 SeitenPIC16F1946madhavaNoch keine Bewertungen
- Low-Power and Area-Efficient Shift Register Using Pulsed Latches With CMOS TechnologyDokument6 SeitenLow-Power and Area-Efficient Shift Register Using Pulsed Latches With CMOS TechnologySesharatnam KoppulaNoch keine Bewertungen
- Cellular Respiration ExplanationDokument2 SeitenCellular Respiration Explanationvestermail50% (2)
- Full Download Strategic Management Text and Cases 9th Edition Dess Test BankDokument35 SeitenFull Download Strategic Management Text and Cases 9th Edition Dess Test Banksaumvirgen2375812100% (28)
- Olympic Message SystemDokument16 SeitenOlympic Message Systemtrevor randyNoch keine Bewertungen
- 36-217 Fall 2013 HW04Dokument2 Seiten36-217 Fall 2013 HW04Gabriel Bernard MullenNoch keine Bewertungen
- Distillation of A Mixture of N-Hexane and N-Heptane-lab - Handout - Experiment - 7Dokument12 SeitenDistillation of A Mixture of N-Hexane and N-Heptane-lab - Handout - Experiment - 7megacobNoch keine Bewertungen
- 2004 Schuhly Major Herbs of Ayurveda E M Williamson Ed Churchill Livingstone New York 2002 361pp Price 49 95 ISBN 0 443 07203 5 HCDokument2 Seiten2004 Schuhly Major Herbs of Ayurveda E M Williamson Ed Churchill Livingstone New York 2002 361pp Price 49 95 ISBN 0 443 07203 5 HCnermal93Noch keine Bewertungen
- Beekman - Tomorrow's Technology and You 9eDokument89 SeitenBeekman - Tomorrow's Technology and You 9eJimmy DrakeNoch keine Bewertungen
- 2nd Year English Send Up 22Dokument2 Seiten2nd Year English Send Up 22Keats 28Noch keine Bewertungen
- Fault Seal Analysis, UK Noerthern Sea, The Alwyn North Statgjord ExampleDokument5 SeitenFault Seal Analysis, UK Noerthern Sea, The Alwyn North Statgjord ExampleAiwarikiaarNoch keine Bewertungen
- H14YS-5A-2301-Reminder - 110423Dokument3 SeitenH14YS-5A-2301-Reminder - 110423BẢO Nhi LêNoch keine Bewertungen
- Renalyn N. Selloga Housekeeping NC IiDokument9 SeitenRenalyn N. Selloga Housekeeping NC IiAlex GinNoch keine Bewertungen
- Incy Wincy SpiderDokument16 SeitenIncy Wincy SpiderOlarteCatalinaNoch keine Bewertungen
- p6 Case Study ModifiedDokument109 Seitenp6 Case Study Modifieddreamboy87100% (2)
- Certified in Production and Inventory Management (CPIM)Dokument8 SeitenCertified in Production and Inventory Management (CPIM)dathient76Noch keine Bewertungen
- Command ReferenceDokument368 SeitenCommand Referenceom007Noch keine Bewertungen
- C Test Questions and Answers PDFDokument15 SeitenC Test Questions and Answers PDFVikas Bakoliya100% (1)
- Caillau, Guillon. S. Ephræm. Opera Omnia. 1832. Volume 7.Dokument540 SeitenCaillau, Guillon. S. Ephræm. Opera Omnia. 1832. Volume 7.Patrologia Latina, Graeca et OrientalisNoch keine Bewertungen
- Polite Form and Verb Stems: Not Being Rude in JapanDokument84 SeitenPolite Form and Verb Stems: Not Being Rude in Japaneulea larkaroNoch keine Bewertungen
- Optimal Voltage RegulatorDokument8 SeitenOptimal Voltage RegulatorARVINDNoch keine Bewertungen
- Nlaa Siam.2010Dokument1 SeiteNlaa Siam.2010YesicaAquinoNoch keine Bewertungen
- Loads Dead Loads Imposed Loads Floor Roof Determining Load Per M and m2 WindDokument58 SeitenLoads Dead Loads Imposed Loads Floor Roof Determining Load Per M and m2 Windwaheedopple3998Noch keine Bewertungen
- Sustainable Architecture WikiDokument9 SeitenSustainable Architecture WikisiddiqueaquibNoch keine Bewertungen
- Public Relations vs. Propaganda: Alexandru BASTIANDokument6 SeitenPublic Relations vs. Propaganda: Alexandru BASTIANFatima AshrafNoch keine Bewertungen
- Optimizing DAX: Improving DAX performance in Microsoft Power BI and Analysis ServicesVon EverandOptimizing DAX: Improving DAX performance in Microsoft Power BI and Analysis ServicesNoch keine Bewertungen
- Dark Data: Why What You Don’t Know MattersVon EverandDark Data: Why What You Don’t Know MattersBewertung: 4.5 von 5 Sternen4.5/5 (3)
- Starting Database Administration: Oracle DBAVon EverandStarting Database Administration: Oracle DBABewertung: 3 von 5 Sternen3/5 (2)
- Blockchain Basics: A Non-Technical Introduction in 25 StepsVon EverandBlockchain Basics: A Non-Technical Introduction in 25 StepsBewertung: 4.5 von 5 Sternen4.5/5 (24)
- Grokking Algorithms: An illustrated guide for programmers and other curious peopleVon EverandGrokking Algorithms: An illustrated guide for programmers and other curious peopleBewertung: 4 von 5 Sternen4/5 (16)
- Relational Database Design and ImplementationVon EverandRelational Database Design and ImplementationBewertung: 4.5 von 5 Sternen4.5/5 (5)
- Fusion Strategy: How Real-Time Data and AI Will Power the Industrial FutureVon EverandFusion Strategy: How Real-Time Data and AI Will Power the Industrial FutureNoch keine Bewertungen
- ITIL 4 : Drive Stakeholder Value: Reference and study guideVon EverandITIL 4 : Drive Stakeholder Value: Reference and study guideNoch keine Bewertungen
- SQL QuickStart Guide: The Simplified Beginner's Guide to Managing, Analyzing, and Manipulating Data With SQLVon EverandSQL QuickStart Guide: The Simplified Beginner's Guide to Managing, Analyzing, and Manipulating Data With SQLBewertung: 4.5 von 5 Sternen4.5/5 (46)
- Business Intelligence Strategy and Big Data Analytics: A General Management PerspectiveVon EverandBusiness Intelligence Strategy and Big Data Analytics: A General Management PerspectiveBewertung: 5 von 5 Sternen5/5 (5)
- Microsoft Access Guide to Success: From Fundamentals to Mastery in Crafting Databases, Optimizing Tasks, & Making Unparalleled Impressions [III EDITION]Von EverandMicrosoft Access Guide to Success: From Fundamentals to Mastery in Crafting Databases, Optimizing Tasks, & Making Unparalleled Impressions [III EDITION]Bewertung: 5 von 5 Sternen5/5 (8)
- The Future of Competitive Strategy: Unleashing the Power of Data and Digital Ecosystems (Management on the Cutting Edge)Von EverandThe Future of Competitive Strategy: Unleashing the Power of Data and Digital Ecosystems (Management on the Cutting Edge)Bewertung: 5 von 5 Sternen5/5 (1)
- Joe Celko's SQL for Smarties: Advanced SQL ProgrammingVon EverandJoe Celko's SQL for Smarties: Advanced SQL ProgrammingBewertung: 3 von 5 Sternen3/5 (1)
- Oracle Database 12c Backup and Recovery Survival GuideVon EverandOracle Database 12c Backup and Recovery Survival GuideBewertung: 4.5 von 5 Sternen4.5/5 (3)
- Modelling Business Information: Entity relationship and class modelling for Business AnalystsVon EverandModelling Business Information: Entity relationship and class modelling for Business AnalystsNoch keine Bewertungen











































































































![Microsoft Access Guide to Success: From Fundamentals to Mastery in Crafting Databases, Optimizing Tasks, & Making Unparalleled Impressions [III EDITION]](https://imgv2-1-f.scribdassets.com/img/word_document/610686937/149x198/9ccfa6158e/1714467780?v=1)








